Understanding Enzyme Kinetics with Peptide Tools: The Role of Peptide Substrates in Quantitative Enzymology
Introduction: Bridging Enzyme Kinetics and Peptide-Based Assays
Enzyme kinetics provides the quantitative framework for understanding how enzymes function, interact with substrates, and respond to inhibitors. It is central to disciplines ranging from biochemistry and pharmacology to drug discovery and systems biology. However, accurate kinetic analysis depends critically on the availability of well-defined substrates.
In this context, peptide substrates have emerged as powerful tools for studying enzyme kinetics. By mimicking key recognition motifs of natural protein substrates while maintaining experimental simplicity, peptide-based systems enable precise measurement of enzymatic parameters such as Km, kcat, and catalytic efficiency (kcat/Km).
This article explores how peptide substrates are used to investigate enzyme kinetics, the principles underlying their design, and how they contribute to reliable and reproducible enzymatic assays.
Fundamentals of Enzyme Kinetics
The Michaelis–Menten Framework
Most enzymatic reactions are described using the Michaelis–Menten model, which relates reaction velocity (v) to substrate concentration ([S]):
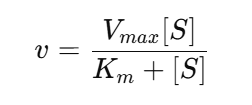
Where:
- Vmax represents the maximum reaction rate
- Km represents the substrate concentration at half-maximal velocity
- kcat (turnover number) describes how many substrate molecules are converted per enzyme per unit time
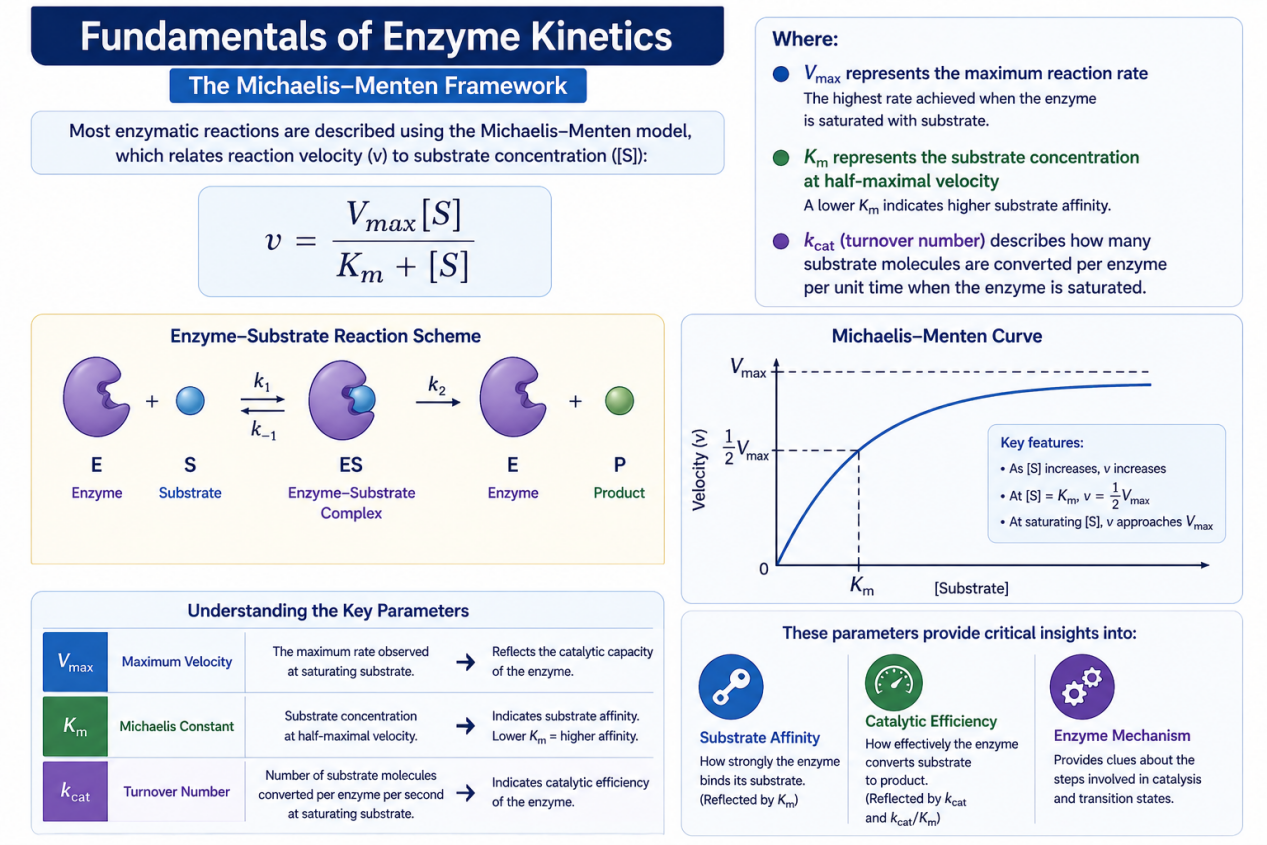
These parameters provide critical insights into:
- Substrate affinity
- Catalytic efficiency
- Enzyme mechanism
Why Substrate Choice Matters
The accuracy of kinetic measurements depends heavily on substrate design. Poor substrate selection can lead to:
- Misleading Km values
- Underestimation of catalytic efficiency
- Non-physiological behavior
Peptide substrates offer a solution by providing controlled, customizable, and reproducible systems.
Why Use Peptide Substrates in Enzyme Kinetics?
Simplified Representation of Complex Systems
Full-length proteins often contain:
- Multiple binding domains
- Complex folding structures
- Allosteric regulation
Peptide substrates isolate the minimal recognition sequence, allowing researchers to focus on the catalytic event itself.
Tunability and Design Flexibility
Peptides can be easily modified to:
- Optimize binding affinity
- Introduce detection labels
- Improve solubility and stability
This flexibility enables systematic investigation of enzyme behavior.
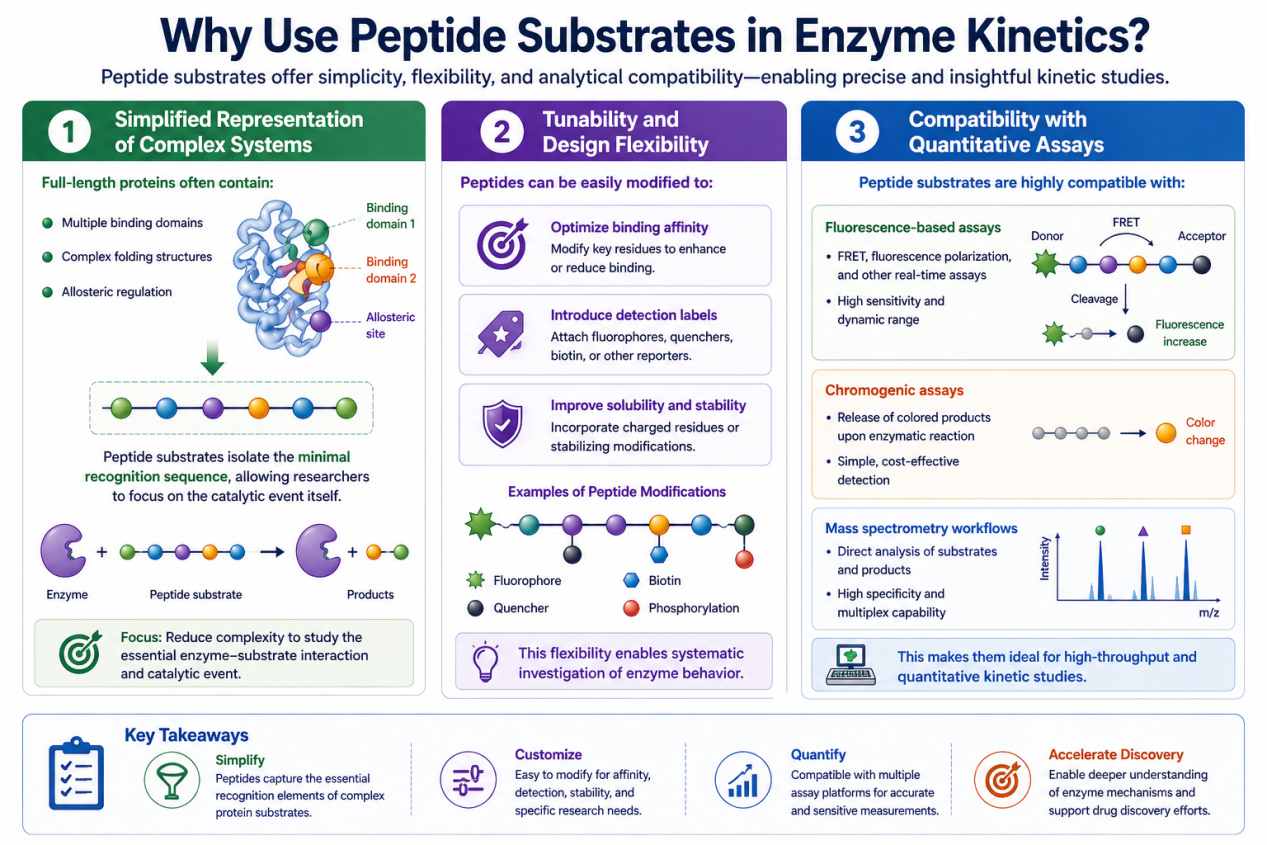
Compatibility with Quantitative Assays
Peptide substrates are highly compatible with:
- Fluorescence-based assays
- Chromogenic assays
- Mass spectrometry workflows
This makes them ideal for high-throughput and quantitative kinetic studies.
Designing Peptide Substrates for Kinetic Analysis
Sequence Specificity and Recognition Motifs
Enzyme–substrate interactions depend on specific amino acid sequences.
Using positional nomenclature:
- P1, P2, P3… represent residues upstream
- P1’, P2’, P3’… represent residues downstream
Each position interacts with corresponding enzyme subsites, influencing:
- Binding affinity
- Catalytic rate
Design Strategy:
- Derive sequences from known physiological substrates
- Optimize key residues through substitution studies
- Validate specificity experimentally
Substrate Length and Structural Context
Peptide length must balance:
- Sufficient binding interactions
- Minimal structural complexity
Considerations:
- Short peptides may lack affinity
- Long peptides may introduce non-specific interactions
In some cases, incorporating secondary structural elements (e.g., β-turns) can enhance activity.
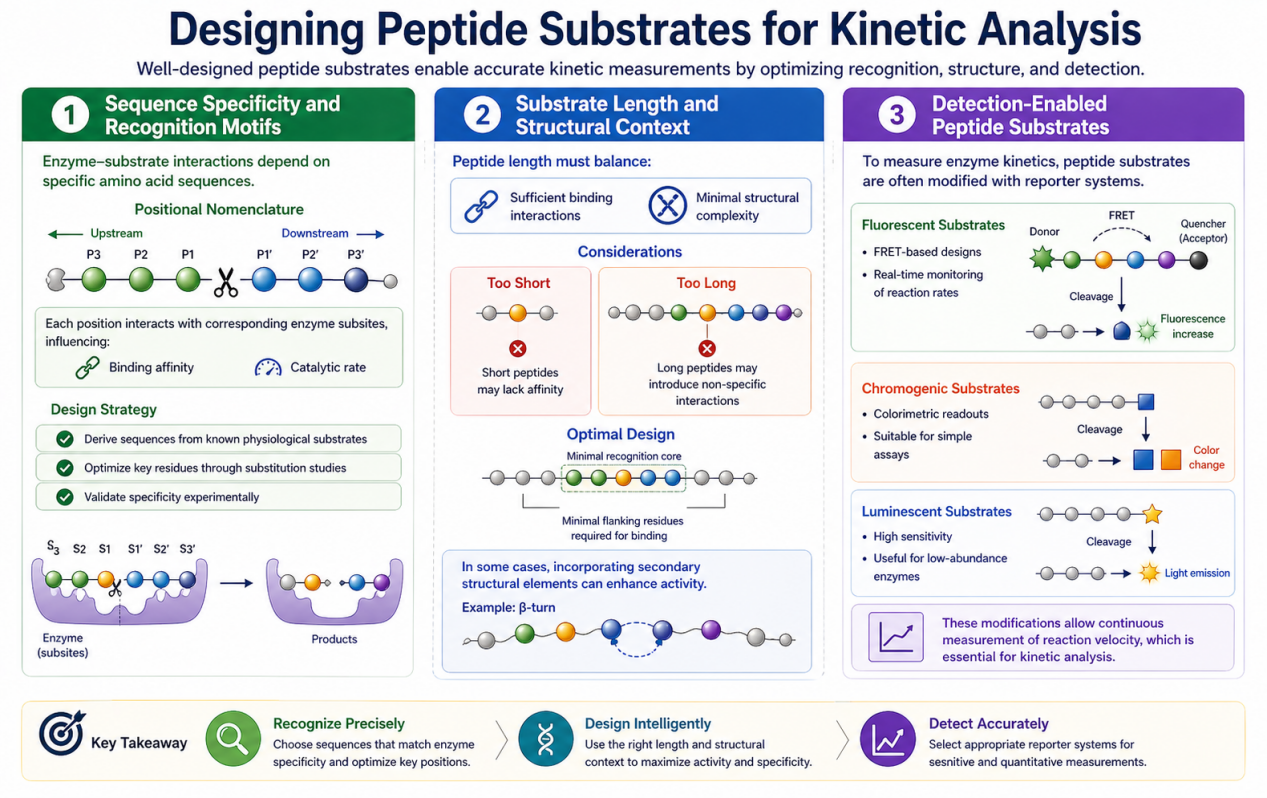
Detection-Enabled Peptide Substrates
To measure enzyme kinetics, peptide substrates are often modified with reporter systems.
Fluorescent Substrates
- FRET-based designs
- Real-time monitoring of reaction rates
Chromogenic Substrates
- Colorimetric readouts
- Suitable for simple assays
Luminescent Substrates
- High sensitivity
- Useful for low-abundance enzymes
These modifications allow continuous measurement of reaction velocity, which is essential for kinetic analysis.
Measuring Kinetic Parameters with Peptide Substrates
Determining Km
Km reflects the affinity between enzyme and substrate.
Using peptide substrates:
- Vary substrate concentration
- Measure initial reaction velocity
- Fit data to the Michaelis–Menten equation
A well-designed peptide substrate should produce:
- A clear saturation curve
- Reliable Km estimation
Determining kcat
kcat represents catalytic turnover.
Calculation requires:
- Known enzyme concentration
- Measured Vmax
Peptide substrates enable accurate determination by:
- Providing consistent reaction conditions
- Minimizing variability
Catalytic Efficiency (kcat/Km)
This parameter reflects overall enzyme performance and is particularly important when:
- Comparing substrates
- Evaluating enzyme mutants
- Screening inhibitors
Advanced Kinetic Applications Using Peptide Tools
Inhibitor Screening
Peptide substrates are widely used in drug discovery to evaluate enzyme inhibitors.
Workflow:
- Measure enzyme activity in presence of inhibitor
- Determine IC50 or Ki values
Peptide substrates allow:
- High-throughput screening
- Quantitative comparison of inhibitor potency
Mechanistic Studies
By modifying peptide substrates, researchers can investigate:
- Transition states
- Catalytic mechanisms
- Substrate recognition patterns
Examples include:
- Substitution of key residues
- Incorporation of non-cleavable analogs
Multiplexed and High-Throughput Assays
Modern assays often use:
- Microplate-based systems
- Automated detection
Peptide substrates are ideal for these applications due to:
- Ease of synthesis
- Consistent performance
Advantages of Peptide-Based Kinetic Assays
High Specificity
Peptide sequences can be tailored to match enzyme preferences, reducing off-target effects.
Reproducibility
Synthetic peptides provide:
- Consistent quality
- Batch-to-batch reproducibility
Flexibility
Peptides can be:
- Modified with labels
- Optimized for different assay conditions
Scalability
Suitable for:
- Small-scale mechanistic studies
- Large-scale screening platforms
Limitations and Considerations
Despite their advantages, peptide substrates have limitations.
Lack of Higher-Order Structure
Peptides may not fully replicate:
- Protein folding
- Allosteric effects
Reduced Biological Complexity
Simplified systems may not capture:
- Cellular context
- Protein–protein interactions
Potential for Non-Physiological Behavior
Over-optimization may lead to:
- Artificially high activity
- Reduced biological relevance
Careful validation is essential.
Best Practices for Using Peptide Substrates in Kinetic Studies
To ensure reliable data:
✔ Use biologically relevant sequences
Maintain connection to natural substrates
✔ Optimize assay conditions
Control pH, temperature, and ionic strength
✔ Validate substrate performance
Confirm specificity and kinetic behavior
✔ Use appropriate controls
Include blank, positive, and negative controls
✔ Perform replicate measurements
Ensure statistical reliability
Emerging Trends in Peptide-Based Kinetics
Recent advances are expanding the role of peptide tools in enzymology:
Integration with Mass Spectrometry
Quantitative MS enables:
- Precise measurement of reaction products
- Multiplexed kinetic analysis
Smart and Activatable Substrates
Peptides that respond dynamically to enzymatic activity:
- Improve sensitivity
- Enable real-time monitoring
Computational Design
Machine learning and modeling are being used to:
- Predict optimal peptide substrates
- Accelerate assay development
How LinkPeptide Supports Enzyme Kinetics Research
At LinkPeptide, we provide tailored solutions for enzyme assay development, including:
- Custom peptide substrate design
- Fluorescent and labeled peptide synthesis
- Sequence optimization for kinetic studies
- High-purity peptides with analytical validation (HPLC, MS)
Our expertise enables researchers to perform accurate, reproducible, and scalable enzyme kinetics studies.
Conclusion
Peptide substrates have become indispensable tools for studying enzyme kinetics, offering a balance between biological relevance and experimental control. By enabling precise measurement of kinetic parameters and supporting advanced assay formats, peptide-based systems play a critical role in modern enzymology.
As analytical technologies and peptide engineering continue to evolve, the integration of peptide tools into kinetic studies will further enhance our understanding of enzyme function and accelerate progress in drug discovery and biomedical research.
Reference
Davidi, D., & Milo, R. (2017). Lessons on enzyme kinetics from quantitative proteomics. Current opinion in biotechnology, 46, 81-89.
https://doi.org/10.1016/j.copbio.2017.02.007
Paradis-Bleau, C., Lloyd, A., Sanschagrin, F., Maaroufi, H., Clarke, T., Blewett, A., … & Levesque, R. C. (2009). Pseudomonas aeruginosa MurE amide ligase: enzyme kinetics and peptide inhibitor. Biochemical Journal, 421(2), 263-272.
https://doi.org/10.1042/BJ20081395
Fu, Y., Alashi, A. M., Young, J. F., Therkildsen, M., & Aluko, R. E. (2017). Enzyme inhibition kinetics and molecular interactions of patatin peptides with angiotensin I-converting enzyme and renin. International journal of biological macromolecules, 101, 207-213.
https://doi.org/10.1016/j.ijbiomac.2017.03.054
Boulware, K. T., Jabaiah, A., & Daugherty, P. S. (2010). Evolutionary optimization of peptide substrates for proteases that exhibit rapid hydrolysis kinetics. Biotechnology and bioengineering, 106(3), 339-346.
 LinkPeptide
LinkPeptide