AI-Driven Design of Antiviral Peptides: A Hybrid Generative–Predictive Framework for Next-Generation Therapeutics
Abstract
The increasing threat of viral pathogens and the limitations of existing antiviral therapeutics underscore the need for more efficient strategies to identify novel antiviral peptides (AVPs). This article examines a recently developed hybrid deep-learning framework that integrates generative modeling with predictive classification to accelerate AVP discovery. Using a Wasserstein GAN with Gradient Penalty (WGAN-GP), the system generates structurally diverse peptide candidates, which are subsequently screened by BiLSTM-based general and virus-specific classifiers trained on curated antiviral datasets. Additional analyses—including structural modeling, motif identification, and drug-likeness filtering—further refine the selection of peptides with strong therapeutic potential. The resulting computational pipeline demonstrates the capacity to explore new regions of peptide space, identify multi-target antiviral candidates, and guide rational peptide engineering. This framework represents a significant advancement toward scalable, AI-driven antiviral peptide discovery, with promising implications for future therapeutic development.
The Rising Need for Accelerated Antiviral Peptide Discovery
Viral pathogens continue to impose a profound global health and economic burden, with infections such as HIV, dengue, influenza, and coronaviruses remaining persistent threats despite decades of therapeutic development. Although antiviral drugs have improved clinical outcomes, their efficacy is often constrained by narrow target specificity, limited availability for emerging pathogens, and the rapid emergence of drug-resistant viral strains. These challenges highlight an urgent need for new classes of broad-spectrum antiviral agents capable of addressing both known and novel viral threats.
Antiviral peptides (AVPs) have emerged as promising therapeutic candidates due to their ability to disrupt viral envelopes, inhibit entry, and modulate host–virus interactions. Yet, progress in AVP discovery remains slow, largely constrained by limited experimental datasets and the high cost of wet-lab screening. As a result, computational approaches—particularly those powered by artificial intelligence—have become increasingly attractive for accelerating peptide design and evaluation.
The study examined in this article introduces a hybrid deep-learning framework that integrates generative modeling with predictive classification, offering a scalable strategy for designing structurally diverse, biologically plausible antiviral peptides.
Building Reliable Antiviral Peptide Datasets: A Computational Starting Point
A critical component of any machine learning framework for peptide discovery is the construction of high-quality datasets that accurately represent both active and inactive molecules. In this study, the authors assembled two complementary databases to support general and virus-specific antiviral peptide modeling. The first, referred to as Database A, integrates more than five thousand experimentally validated antiviral peptides alongside a curated set of non-AVPs. This binary dataset enables the development of a broad classifier capable of distinguishing peptides with putative antiviral properties from those lacking documented activity.
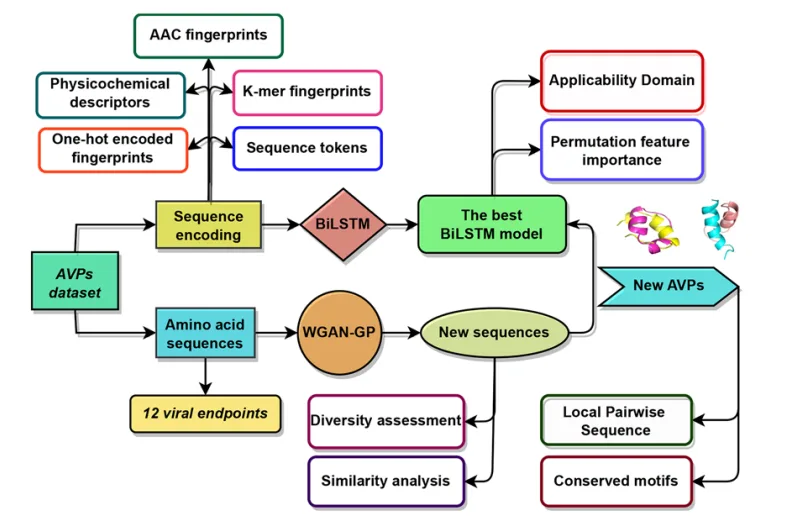
To further refine predictions toward particular pathogens, the authors constructed Database B, a collection of virus-specific AVPs covering twelve clinically relevant enveloped viruses. Unlike the generalized dataset, Database B captures the diversity of antiviral mechanisms and sequence patterns associated with individual viral families, allowing for the development of specialized predictive models.
Feature engineering represents another foundational aspect of the pipeline. By combining amino acid composition, K-mer fingerprints, one-hot encodings, physicochemical descriptors, and tokenized sequence representations, the authors ensure that both structural and biochemical information is encoded for downstream learning. This multi-feature strategy provides a robust foundation for training deep-learning models capable of capturing the complex determinants of antiviral activity.
Learning the Language of Antiviral Peptides: A BiLSTM-Based Prediction Model
While predictive models are effective for classifying known peptides, they cannot independently expand the chemical space of antiviral candidates. To address this limitation, the study integrates a generative deep-learning component based on the Wasserstein Generative Adversarial Network with Gradient Penalty (WGAN-GP). This architecture offers significant advantages over conventional GANs, particularly in stabilizing training dynamics and mitigating mode collapse—issues that are especially problematic when working with small or heterogeneous biological datasets.
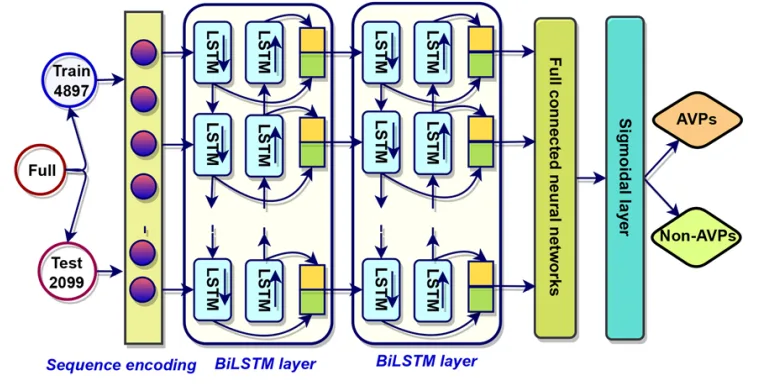
The generator network receives random noise vectors and transforms them into candidate peptide sequences encoded in one-hot format, while the critic network evaluates how closely these generated sequences resemble experimentally validated AVPs. Through iterative adversarial training, the generator learns to produce peptides that increasingly align with the statistical and structural distributions of known antiviral sequences.
Robust validation analyses confirm the biological plausibility of the generated peptides. Dimensionality reduction techniques such as t-SNE and UMAP demonstrate a substantial overlap between generated and real AVPs, indicating that the model successfully captures the underlying sequence space. Additionally, diversity metrics—including Shannon entropy and pairwise sequence variation—show that the generative model preserves essential variability while exploring novel regions of peptide space.
Expanding Peptide Chemical Space with WGAN-GP–Driven Design
While predictive models are effective for classifying known peptides, they cannot independently expand the chemical space of antiviral candidates. To address this limitation, the study integrates a generative deep-learning component based on the Wasserstein Generative Adversarial Network with Gradient Penalty (WGAN-GP). This architecture offers significant advantages over conventional GANs, particularly in stabilizing training dynamics and mitigating mode collapse—issues that are especially problematic when working with small or heterogeneous biological datasets.
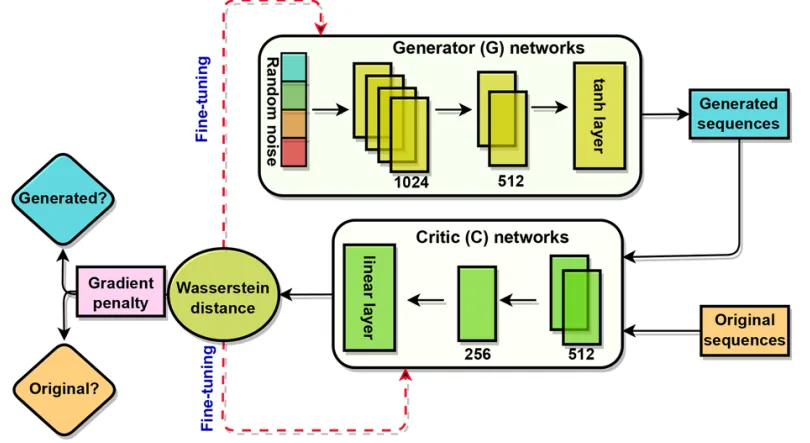
The generator network receives random noise vectors and transforms them into candidate peptide sequences encoded in one-hot format, while the critic network evaluates how closely these generated sequences resemble experimentally validated AVPs. Through iterative adversarial training, the generator learns to produce peptides that increasingly align with the statistical and structural distributions of known antiviral sequences.
Robust validation analyses confirm the biological plausibility of the generated peptides. Dimensionality reduction techniques such as t-SNE and UMAP demonstrate a substantial overlap between generated and real AVPs, indicating that the model successfully captures the underlying sequence space. Additionally, diversity metrics—including Shannon entropy and pairwise sequence variation—show that the generative model preserves essential variability while exploring novel regions of peptide space.
A Unified AI Pipeline for High-Throughput Antiviral Peptide Discovery
The most innovative aspect of the study lies in its hybrid workflow, which integrates generative modeling with predictive classification to form a unified antiviral peptide discovery pipeline. In the first stage, the WGAN-GP model produces a library of novel peptide sequences that mirror the statistical properties of known AVPs while extending into unexplored regions of sequence space. These generated peptides are then subjected to a rigorous second-stage evaluation using both the general BiLSTM classifier and virus-specific predictive models.
This two-step screening strategy enables efficient prioritization of candidates with high antiviral potential. From an initial set of 300 generated peptides, the authors identified 815 antiviral “hits” across 12 enveloped viruses, reflecting the model’s capacity to propose peptides with multi-target potential—a desirable characteristic for broad-spectrum antiviral development.
To further assess the structural plausibility of the top candidates, representative peptides were modeled using AlphaFold3, revealing diverse secondary structure features consistent with known antiviral mechanisms. The integration of sequence generation, predictive scoring, and structural modeling thus provides a streamlined computational framework for advancing peptide candidates from concept to preclinical prioritization.
Toward AI-Guided Development of Next-Generation Antiviral Peptides
This study demonstrates the transformative potential of integrating generative and predictive deep-learning frameworks to accelerate antiviral peptide discovery. By combining WGAN-GP–driven sequence generation with highly discriminative BiLSTM classifiers, the authors present a scalable workflow capable of producing structurally diverse, biologically plausible AVP candidates. The subsequent incorporation of structural prediction, motif analysis, and drug-likeness filtering further strengthens the translational relevance of the proposed pipeline.
Importantly, the work highlights how AI can address longstanding challenges in peptide therapeutics, including limited dataset availability, uneven viral representation, and the high cost of experimental screening. Although experimental validation remains essential, the hybrid methodology offers a powerful foundation for rapidly exploring antiviral peptide space and prioritizing candidates for laboratory investigation. As viral threats continue to evolve, such computational approaches will play an increasingly critical role in guiding the design of next-generation antiviral agents.
Reference
Duy, H. A., & Srisongkram, T. (2025). A hybrid framework of generative deep learning for antiviral peptide discovery. Scientific Reports, 15(1), 25554.https://doi.org/10.1038/s41598-025-11328-9
Murray, C. J., Aravkin, A. Y., Zheng, P., Abbafati, C., Abbas, K. M., Abbasi-Kangevari, M., … & Borzouei, S. (2020). Global burden of 87 risk factors in 204 countries and territories, 1990–2019: a systematic analysis for the Global Burden of Disease Study 2019. The lancet, 396(10258), 1223-1249.https://doi.org/10.1016/S0140-6736(20)30752-2
Rossino, G., Marchese, E., Galli, G., Verde, F., Finizio, M., Serra, M., … & Collina, S. (2023). Peptides as therapeutic agents: challenges and opportunities in the green transition era. Molecules, 28(20), 7165.https://doi.org/10.3390/molecules28207165
Surana, S., Arora, P., Singh, D., Sahasrabuddhe, D., & Valadi, J. (2023). Pandoragan: generating antiviral peptides using generative adversarial network. SN Computer Science, 4(5), 607.https://doi.org/10.1007/s42979-023-02203-3
 LinkPeptide
LinkPeptide