From Bottlenecks to Breakthroughs: Redesigning Microbial Strains for Enhanced Peptide Production
Abstract
Non-ribosomal peptides (NRPs) are a diverse class of bioactive compounds with immense pharmaceutical and industrial value. However, their biosynthesis is often limited by complex regulatory networks in native microbial hosts, hindering scalable production. This blog explores a recent study that introduces a rational, multi-target strategy to enhance NRP yield through genome-wide CRISPR interference (CRISPRi) and a synergy-guided combination framework. A novel colorimetric reporter system using indigoidine enables efficient screening of high-yield phenotypes. By integrating this with CRISPRi libraries and synergy coefficient analysis, the researchers identified and combined cooperative genetic repressors to dramatically increase titers of daptomycin, thaxtomin A, and surfactin in Streptomyces and Bacillus species. This strategy not only sets a new benchmark for strain engineering but also provides a scalable model for optimizing secondary metabolite production across diverse microbial platforms.
Overcoming the Regulatory Complexity of Non-Ribosomal Peptide Production
Non-ribosomal peptides (NRPs) represent a diverse and highly valuable class of secondary metabolites with extensive pharmaceutical and agricultural applications. These compounds—produced independently of the ribosomal translation machinery—include prominent antibiotics such as daptomycin, antitumor agents, immunosuppressants, and biosurfactants like surfactin. They are typically synthesized by large, modular non-ribosomal peptide synthetases (NRPSs) encoded within biosynthetic gene clusters (BGCs), predominantly in Streptomyces and Bacillus species.
Despite their significance, the industrial production of NRPs remains a considerable challenge. Native strains often exhibit low yields due to complex, multi-layered regulatory networks governing BGC expression. These networks are evolutionarily optimized for survival under specific environmental conditions rather than for maximizing metabolite output. Traditional approaches to enhance NRP production, such as mutagenesis and overexpression of pathway-specific regulators, are largely empirical and frequently result in suboptimal or unpredictable outcomes.
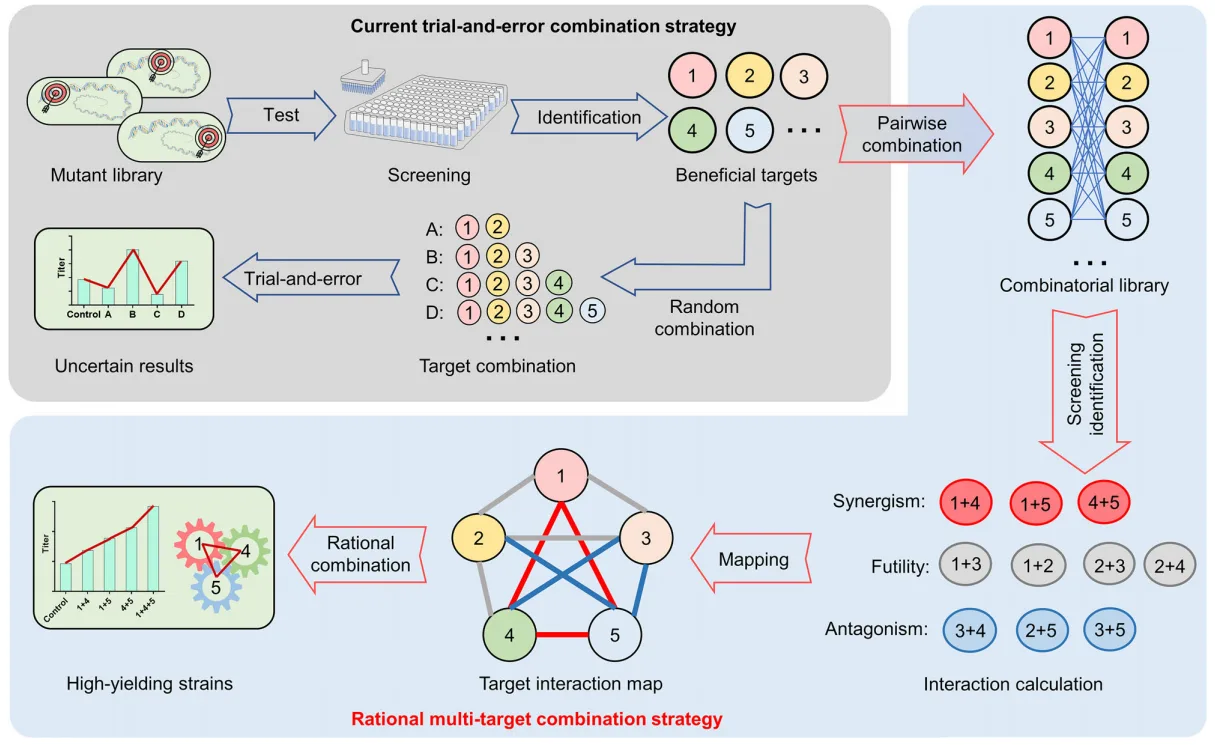
To overcome these limitations, the study by Yan et al. introduces a rational, multi-target strategy that leverages CRISPR interference (CRISPRi) to systematically perturb gene regulation at the genome scale. By integrating a novel colorimetric reporter system and a synergy-guided combination framework, the authors present a comprehensive methodology to identify and combine multiple beneficial regulatory targets, thereby achieving a substantial and reproducible improvement in NRP biosynthesis.
A Novel Reporter System for High-Throughput Phenotype Screening
A major limitation in high-throughput strain engineering for non-ribosomal peptide (NRP) production is the lack of robust, scalable, and universal screening systems. Conventional methods often rely on indirect indicators such as growth-coupled phenotypes, riboswitch-based biosensors, or labor-intensive metabolite quantification, which are either insufficiently sensitive or poorly applicable across diverse biosynthetic systems. To address this gap, the authors of the study devised an innovative analog co-expression and co-biosynthesis reporter platform that enables real-time, colorimetric assessment of NRP production potential.
Central to this system is the heterologous expression of the indigoidine biosynthetic gene cluster, a minimal two-gene module producing a distinctive blue pigment. By integrating the idgS-sfp cassette downstream of the dptD gene in the daptomycin biosynthetic gene cluster (BGC) of Streptomyces roseosporus, the authors ensured that indigoidine synthesis would be co-regulated with native NRP production. This design facilitates synchronized transcription and translation, effectively coupling the output of daptomycin with that of the easily measurable indigoidine pigment.
Experimental validation confirmed a strong positive correlation between indigoidine intensity and daptomycin titers, demonstrating that indigoidine can serve as a faithful proxy for production phenotypes. Importantly, this system captured not only changes in BGC expression but also broader physiological and regulatory alterations affecting biosynthetic capacity. The simplicity, modularity, and transferability of this reporter approach mark a significant advancement over previous biosensor technologies, enabling rapid, reliable screening of high-yield phenotypes in both homologous and heterologous hosts.
CRISPRi-Driven Genome-Wide Screening and Synergistic Target Mapping
To systematically identify genetic determinants that suppress non-ribosomal peptide (NRP) biosynthesis, the authors employed a genome-scale CRISPR interference (CRISPRi) platform in Streptomyces roseosporus. By targeting 610 predicted regulatory genes using a comprehensive library of single-guide RNAs (sgRNAs), each designed to inhibit transcription by targeting the non-template DNA strand, the study enabled high-throughput repression of regulatory elements across the genome. This approach was integrated with the previously established indigoidine-based reporter system, allowing for rapid visual screening of mutants with enhanced NRP biosynthetic potential.
Through this strategy, the authors identified 17 regulatory genes whose repression led to significant increases in daptomycin production, each showing between 17–38% improvement in yield relative to control strains. These genes, designated idt1 through idt17, encompass a diverse array of transcription factor families, including TetR, MarR, and GntR, many of which were previously uncharacterized in the context of secondary metabolism. This underscores the strength of unbiased, large-scale screening in uncovering novel regulatory levers.
Critically, the study advanced beyond single-target engineering by exploring combinatorial effects between identified repressors. To do so, the authors developed a dual-sgRNA CRISPRi system capable of simultaneously silencing gene pairs. The combinatorial effects were quantified using a synergy coefficient (q), adapted from pharmacological models of drug interaction. Combinations were classified as antagonistic (q < 0.85), additive (0.85 ≤ q < 1.15), or synergistic (q ≥ 1.15), enabling the construction of an interaction network that revealed a cluster of four regulators—idt3, idt5, idt9, and idt15—with consistently positive synergistic effects.
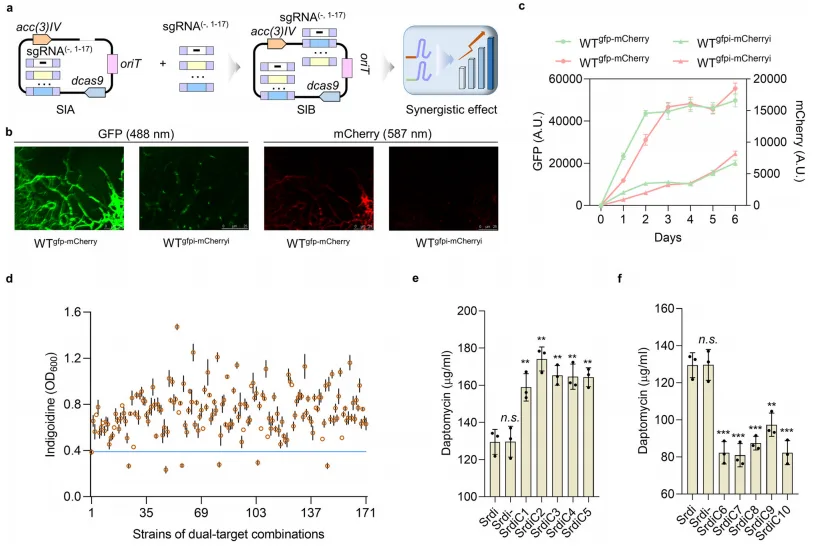
Engineering a strain (termed Sr11) with simultaneous repression of these four targets yielded a dramatic increase in daptomycin titer, reaching 1054 mg/L in a 7.5-L fermenter—doubling the previous maximum reported yield. This result not only validates the utility of synergy-guided combinatorial engineering but also sets a new benchmark for industrial daptomycin production.
Expanding the Strategy Across Hosts and Peptide Classes
One of the most compelling aspects of the study lies in the successful generalization of the multi-target rational engineering framework beyond the original host species. To assess the broader applicability of their strategy, the authors extended their workflow to two additional microbial systems: Streptomyces coelicolor for the production of thaxtomin A, and Bacillus subtilis for the biosynthesis of surfactin. Both compounds—thaxtomin A, a plant-derived herbicidal agent, and surfactin, a potent cyclic lipopeptide biosurfactant—are representative of the structural and functional diversity of non-ribosomal peptides (NRPs).
In S. coelicolor, the thaxtomin biosynthetic gene cluster (BGC), introduced heterologously, exhibited cryptic expression under standard conditions. By implementing the analog co-expression and co-biosynthesis reporter system, now adapted to track thaxtomin production via indigoidine levels, the authors screened a CRISPRi library targeting 856 regulatory genes. This led to the identification of 10 repressor genes whose inhibition reactivated the silent BGC. Subsequent combinatorial analysis and synergy mapping revealed that the triple-target combination of thaR4, thaR7, and thaR8 exhibited strong positive interactions, resulting in a high-yielding strain producing thaxtomin A at 352 mg/L in shake flasks—a significant advance in heterologous expression systems.
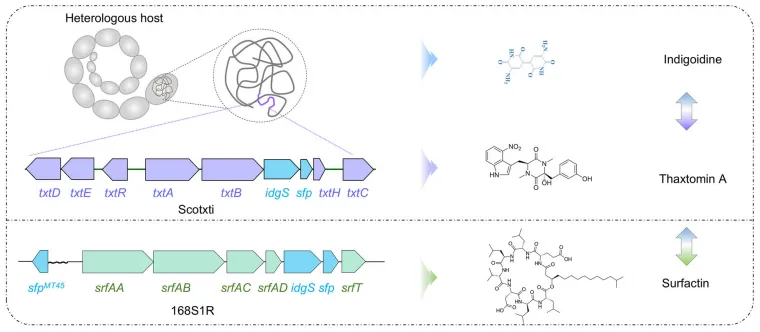
A similar approach was applied to B. subtilis 168, wherein surfactin biosynthesis was initially activated by introducing a heterologous sfp gene. CRISPRi screening identified 11 repressors affecting surfactin production, with synergy-guided engineering of srfR1, srfR5, srfR7, and srfR10 culminating in a strain that produced 878 mg/L of surfactin—nearly four times the baseline level.
These results collectively demonstrate that the described strategy is not confined to a single species or NRP class. Rather, it constitutes a broadly applicable toolkit for activating silent biosynthetic pathways and optimizing yield across genetically and physiologically diverse microbial hosts.
Redefining Strain Engineering Through Rational Multi-Target Design
The study by Yan et al. represents a transformative shift in the way microbial strains are engineered for high-value metabolite production. Traditional strategies—such as random mutagenesis, pathway-specific overexpression, or empirical target combinations—often rely on labor-intensive trial-and-error and yield unpredictable or suboptimal outcomes. These approaches rarely account for the complex, non-linear interactions between genetic elements that govern secondary metabolism.
In contrast, the rational multi-target combination strategy developed in this work offers a data-driven, systematic framework grounded in genome-scale screening and quantitative synergy mapping. By incorporating CRISPRi-mediated gene perturbation, an indigoidine-based phenotypic reporter, and a synergy coefficient model, the approach enables the identification and integration of truly cooperative genetic interventions. This significantly accelerates strain development, reduces experimental burden, and enhances reproducibility across diverse microbial systems.
The broader implications of this study are substantial. In synthetic biology and industrial biotechnology, such strategies open new avenues for unlocking silent biosynthetic gene clusters, optimizing heterologous production, and constructing robust microbial cell factories. The integration of high-throughput genetic tools with network-level insights offers a scalable blueprint for precision metabolic engineering—poised to impact drug discovery, agricultural biotechnology, and the sustainable manufacturing of complex natural products.
 LinkPeptide
LinkPeptide