From Code to Cure: How AfCycDesign is Revolutionizing Cyclic Peptide Therapeutics
Abstract
Cyclic peptides hold immense promise in therapeutic development, but their complex structures have made accurate design challenging. AfCycDesign, a novel deep learning framework built on AlphaFold2, overcomes these limitations by enabling reliable structure prediction, sequence redesign, and de novo generation of cyclic peptides. This approach achieves atomic-level accuracy with high efficiency, validated by multiple X-ray structures and functional peptide binders targeting proteins like MDM2 and Keap1. AfCycDesign not only accelerates peptide drug discovery but also opens new possibilities for targeting previously inaccessible biological interactions, marking a significant advancement in the field of computational peptide design.
Old Challenges, New Shapes: The Promise and Complexity of Cyclic Peptides
Cyclic peptides are rapidly gaining traction in drug discovery, celebrated for their exceptional structural stability, resistance to enzymatic degradation, and unique ability to modulate protein–protein interactions—a class of targets often considered “undruggable” by small molecules or antibodies. Their constrained ring-like architecture locks them into defined shapes, allowing for high specificity and affinity in therapeutic applications ranging from oncology to infectious diseases.
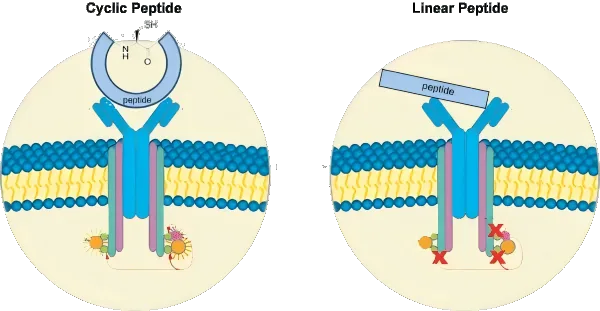
However, this very rigidity presents a major hurdle: designing cyclic peptides computationally is notoriously difficult. Traditional modeling tools struggle to predict their diverse topologies and folding patterns, largely due to a lack of high-resolution structural data and the challenges of capturing their closed-loop geometry.
While deep learning tools like AlphaFold2 have transformed protein structure prediction, they weren’t built with cyclic peptides in mind. This has left a critical gap in peptide therapeutics—until now.
Enter AfCycDesign, a powerful new framework that adapts AlphaFold2’s architecture for the specific challenges of cyclic peptide modeling. With this innovation, researchers can now design, predict, and optimize cyclic peptides with unprecedented accuracy—paving the way for a new generation of peptide-based therapeutics.
Closing the Loop: How AfCycDesign Adapts AlphaFold2 for Cyclic Peptides
At the heart of AfCycDesign is a deceptively simple idea: what if AlphaFold2, the deep learning model that reshaped protein structure prediction, could be adapted to understand cyclic peptides?
AlphaFold2 was originally designed for linear proteins, using positional encoding to represent distances between residues in a sequence. But cyclic peptides—lacking free N- and C-termini—require a different logic. To address this, the AfCycDesign team engineered a cyclic offset matrix that rewires AlphaFold2’s perception of the peptide chain, effectively “closing the loop” during structure prediction.
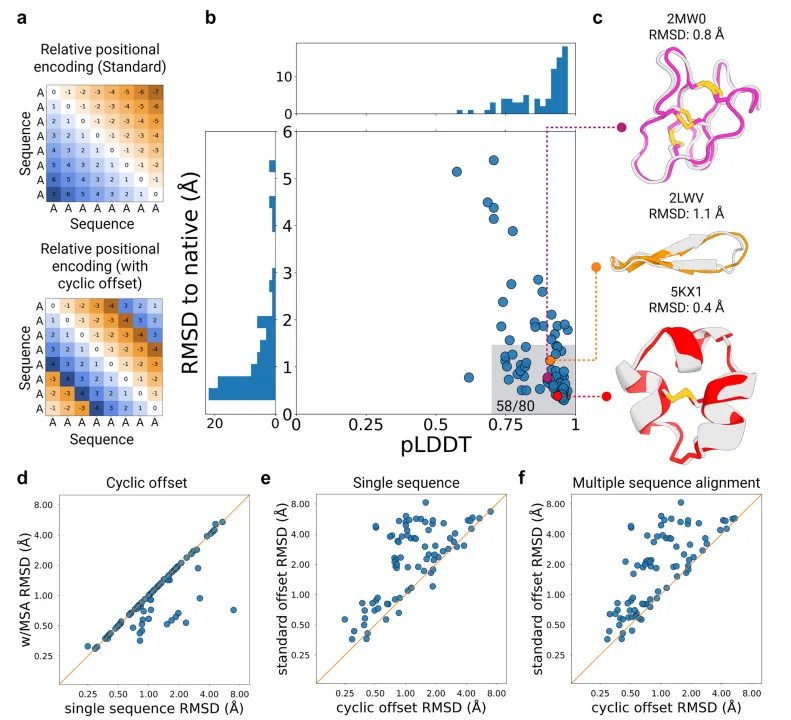
Structure prediction of native cyclic peptides using AfCycDesign
This innovation proved transformative. In benchmarking tests, AfCycDesign was able to accurately predict the 3D structures of 58 out of 80 cyclic peptides from the Protein Data Bank, with RMSD values below 1.5 Å and confidence scores (pLDDT) above 0.7. Even more impressively, 80% of high-confidence predictions (pLDDT > 0.85) closely matched the experimentally determined structures.
Unlike previous physics-based methods like Rosetta, which require intensive sampling and hours of compute time, AfCycDesign delivers predictions in just minutes on a single GPU. It also performs well even without multiple sequence alignments, relying on single-sequence inputs—a critical feature for peptides that often lack deep evolutionary history.
AfCycDesign doesn’t just predict structure—it enables fast, scalable, and accurate exploration of peptide space. It bridges the gap between deep learning and peptide therapeutics, offering researchers a tool to move from idea to atomically precise structure in record time.
Hallucinating Peptides: Creating Entirely New Structures from Scratch
One of the most powerful features of AfCycDesign is its ability to generate entirely new cyclic peptide structures from scratch—a process the authors call hallucination. Unlike redesign, which starts with a known backbone, hallucination involves simultaneous generation of both the sequence and structure, guided by internal metrics that reward compactness, confidence (pLDDT), and low alignment error (PAE).
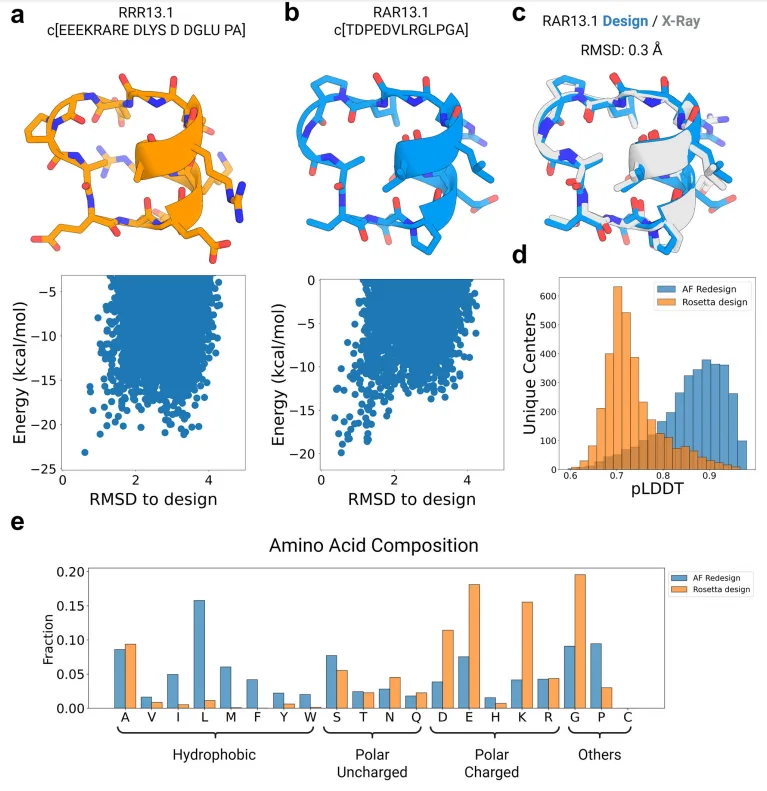
Sequence design of cyclic peptide backbones using AfCycDesign
To test this capability, the researchers launched large-scale hallucination runs across peptides ranging from 7 to 13 residues. The result was staggering: more than 10,000 unique structural clusters were identified, many of which passed strict confidence thresholds (pLDDT > 0.9). These designs spanned a wide variety of topologies—compact loops, turns, beta strands, and even short helices—capturing a rich and previously unexplored design space.
Critically, these predictions weren’t just theoretical. The team synthesized and crystallized eight of these hallucinated peptides. Each structure, from the smallest 7-mer (RH7.1) to the more complex 13-mer (RH13.1), showed striking agreement with the design models, achieving Cα RMSDs below 1.0 Å. This level of precision, previously elusive in de novo peptide design, validates AfCycDesign’s power to create accurate, foldable scaffolds from scratch.
These high-confidence scaffolds serve as ready-to-use templates for downstream applications. Whether used as carriers, therapeutic agents, or targeting modules, the hallucinated peptides offer a robust and scalable foundation for functionalization—something rarely possible at this level of accuracy and diversity.
Designed to Bind: Building Functional Peptide Inhibitors with Atomic Precision
Designing a well-folded peptide is one thing—but turning it into a therapeutic binder is the real test of functional utility. AfCycDesign clears this bar with ease.
To explore its potential for drug discovery, the researchers used their high-confidence hallucinated scaffolds to design macrocyclic inhibitors targeting two biologically important proteins: MDM2, a key regulator of p53 in cancer, and Keap1, which controls oxidative stress responses through Nrf2 degradation.
For MDM2, the team grafted a known 5-residue p53-derived binding motif (FSDLW) onto hallucinated scaffolds and redesigned the surrounding sequence using ProteinMPNN and Rosetta. From just 16 candidates synthesized, 5 showed strong inhibition in vitro, and 3 exhibited nanomolar IC50 values, with the best (RMG_14) reaching 338 nM. Even more impressively, the X-ray structure of the MDM2–RMG_14 complex matched the predicted model with an RMSD of just 1.0 Å, confirming not only the peptide’s activity but also its precisely engineered binding pose.
The success didn’t stop there. The team applied the same strategy to Keap1, grafting a 4-residue “hot loop” (EETG) from Nrf2 onto new scaffolds. Three of the six synthesized peptides performed on par or better than the native Nrf2 peptide, with KC4 standing out as twice as potent, thanks to a stabilizing hydrogen bond added during design.
These examples underscore AfCycDesign’s unique ability to move from scaffold generation to functional inhibitors—rapidly and with atomic-level control. It’s a potent demonstration of how AI-driven peptide design is poised to transform therapeutic discovery.
A New Era Begins: Intelligent Design for the Next Generation of Peptide Therapeutics
AfCycDesign marks a paradigm shift in peptide design, bridging deep learning with the intricacies of macrocyclic structure. By adapting AlphaFold2 with cyclic-aware positional encoding, it unlocks reliable structure prediction, intelligent sequence redesign, and high-throughput de novo generation—all tailored to the unique demands of cyclic peptides.
The platform’s success goes beyond modeling. From the structural validation of hallucinated peptides to the creation of potent, nanomolar-range binders for MDM2 and Keap1, AfCycDesign demonstrates real translational potential. Its ability to rapidly generate functional peptide scaffolds could accelerate the development of therapeutics for cancer, inflammatory diseases, and beyond.
As researchers explore modifications with D-amino acids and non-canonical residues, the scaffold set generated by AfCycDesign provides a launchpad for innovation. In the coming years, it could redefine how we approach drug targets once thought unreachable—ushering in a new era of programmable peptide medicines.
 LinkPeptide
LinkPeptide