Labeled Peptides: Advances in Quantitative Peptide Analysis
Introduction: The Growing Demand for Quantitative Precision in Peptide Research
In modern biomedical research, the ability to accurately quantify peptides has become increasingly critical. From biomarker discovery and drug development to systems biology and clinical diagnostics, researchers rely on precise measurements to generate reproducible and meaningful data.
However, peptide quantification presents unique challenges. Peptides often exist in low abundance, are prone to enzymatic degradation, and must be analyzed within highly complex biological matrices such as plasma, tissue extracts, or cell lysates. These factors make traditional analytical approaches insufficient for achieving the required level of precision.
To address these limitations, labeled peptides have emerged as indispensable tools in quantitative workflows. By incorporating stable isotopes or chemical tags, labeled peptides enable accurate detection, normalization, and comparison across samples—significantly improving analytical reliability.
Recent advances in mass spectrometry (MS), labeling chemistry, and data analysis have further enhanced the capabilities of labeled peptide strategies, transforming quantitative peptide analysis into a highly sensitive and scalable discipline.
What Are Labeled Peptides in Quantitative Analysis?
In the context of quantitative research, labeled peptides are peptides that have been chemically modified with detectable markers, most commonly:
- Stable isotopes (e.g., 13C, 15N, 2H)
- Isobaric chemical tags (e.g., TMT, iTRAQ)
These modifications introduce mass or signal differences that can be distinguished using analytical techniques such as LC-MS/MS.
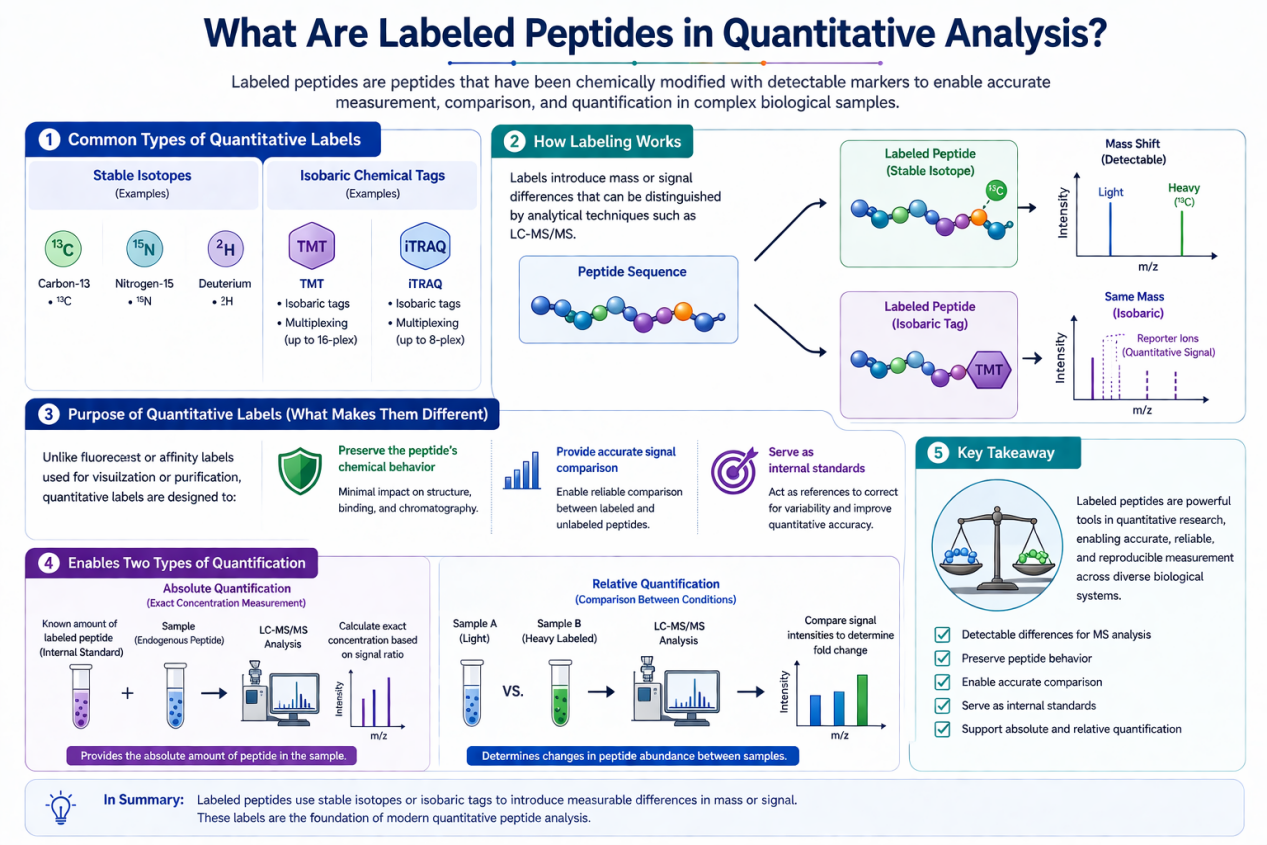
Unlike fluorescent or affinity labels used for visualization or purification, quantitative labels are designed to:
- Preserve the peptide’s chemical behavior
- Provide accurate signal comparison
- Serve as internal standards
This enables researchers to perform both:
- Absolute quantification (exact concentration measurement)
- Relative quantification (comparison between experimental conditions)
Stable Isotope Labeling: The Foundation of Quantitative Peptide Analysis
Stable isotope labeling is widely regarded as the gold standard for quantitative peptide analysis due to its accuracy and minimal interference with peptide function.
Key Principle
Stable isotopes are chemically identical to their natural counterparts but differ in atomic mass. When incorporated into peptides, they produce a predictable mass shift without altering:
- Binding properties
- Chromatographic behavior
- Biological activity
This allows labeled and unlabeled peptides to behave identically during analysis while remaining distinguishable in mass spectrometry.
Absolute Quantification Using the AQUA Strategy
One of the most widely used methods is the AQUA (Absolute QUAntification) approach.
Workflow:
- A synthetic peptide labeled with stable isotopes is prepared at a known concentration
- The labeled peptide is spiked into the biological sample
- LC-MS/MS is used to analyze both labeled and endogenous peptides
- Signal intensities are compared to calculate absolute concentrations
Advantages:
- High precision
- Direct quantification
- Minimal experimental variability
- This approach is particularly valuable in:
- Clinical biomarker validation
- Pharmacokinetic studies
- Targeted proteomics
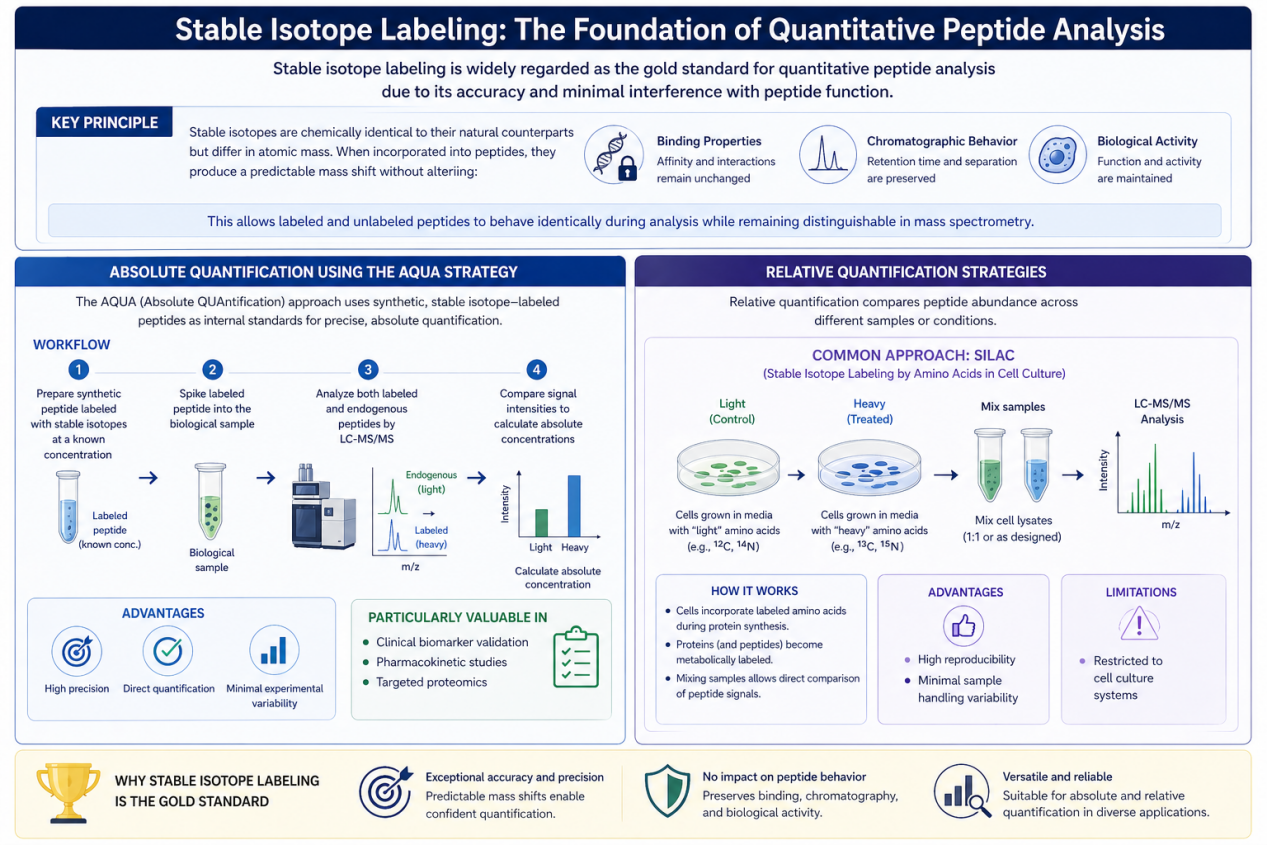

Relative Quantification Strategies
Relative quantification compares peptide abundance across different samples or conditions.
Common Approaches:
SILAC (Stable Isotope Labeling by Amino Acids in Cell Culture)
- Cells are grown in media containing labeled amino acids
- Proteins are metabolically labeled during synthesis
- Enables comparison between treated and control samples
Advantages:
- High reproducibility
- Minimal sample handling variability
Limitations:
- Restricted to cell culture systems
Chemical Labeling Approaches for Multiplexed Analysis
While isotope labeling excels in accuracy, chemical labeling techniques offer advantages in throughput and scalability.
Tandem Mass Tags (TMT)
TMT is an isobaric labeling method that allows simultaneous analysis of multiple samples.
Key Features:
- Tags have identical mass but release different reporter ions during MS/MS
- Enables multiplexing (up to 16 samples or more)
Advantages:
- High throughput
- Reduced experimental variability
- Efficient comparison across conditions
Challenges:
- Ratio compression due to co-isolation
- Requires advanced MS instrumentation
iTRAQ Labeling
iTRAQ operates on a similar principle to TMT, offering:
- Multiplex quantification
- High sensitivity
- Compatibility with complex samples
- Both TMT and iTRAQ are widely used in large-scale proteomics studies.
Advances in Mass Spectrometry Technologies
The effectiveness of labeled peptide strategies is closely tied to advances in mass spectrometry (MS).
High-Resolution Mass Spectrometry
Modern instruments provide:
- Enhanced sensitivity
- Improved mass accuracy
- Detection of low-abundance peptides
This allows researchers to quantify peptides in previously inaccessible biological contexts.
Parallel Reaction Monitoring (PRM)
PRM is a targeted MS technique that improves:
- Quantification accuracy
- Signal specificity
- Reproducibility
Unlike traditional methods, PRM monitors predefined peptide fragments, reducing interference and improving confidence in results.
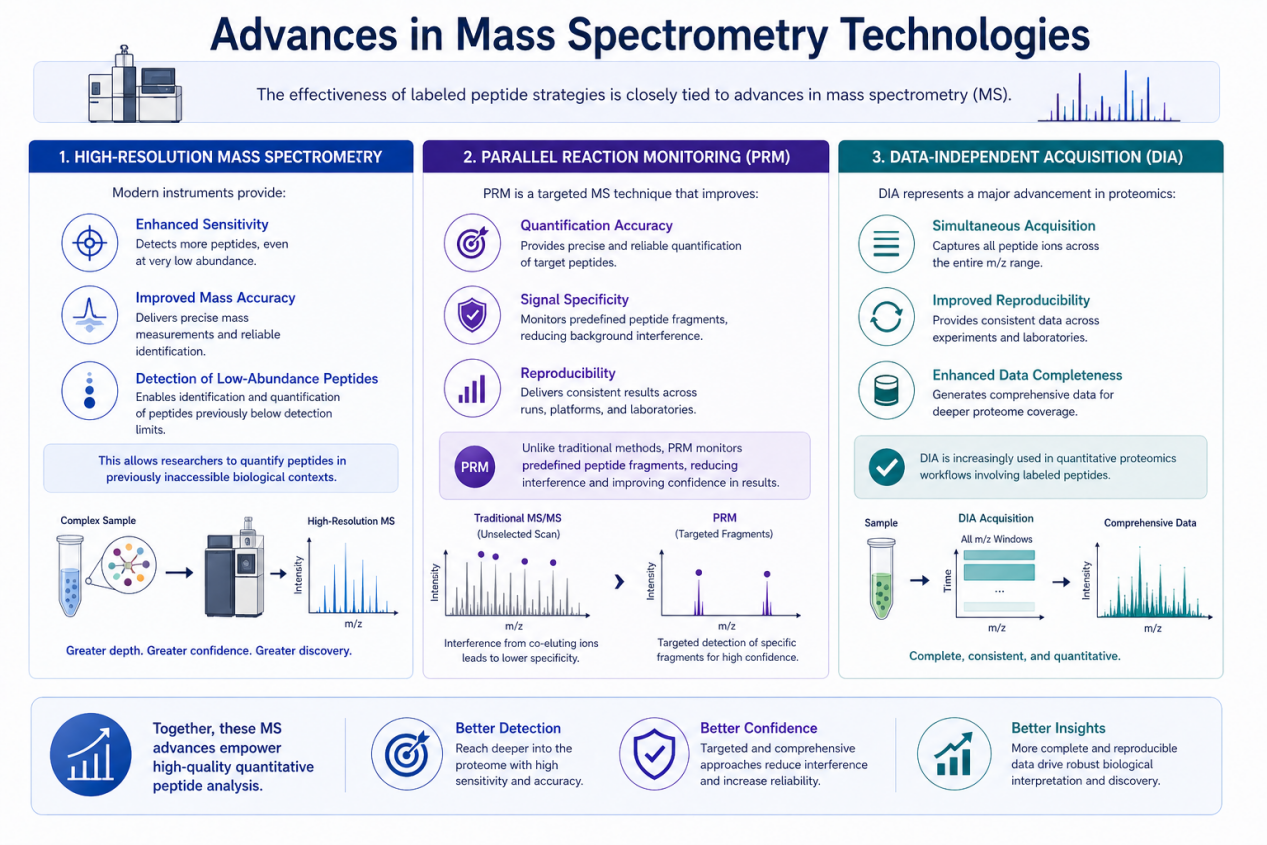
Data-Independent Acquisition (DIA)
DIA represents a major advancement in proteomics:
- Simultaneous acquisition of all peptide ions
- Improved reproducibility across experiments
- Enhanced data completeness
- DIA is increasingly used in quantitative proteomics workflows involving labeled peptides.
Advantages of Labeled Peptides in Quantitative Workflows
Precision and Accuracy
Labeled peptides serve as internal references, minimizing variability caused by:
- Sample preparation
- Instrument fluctuations
- Matrix effects
Reproducibility
Standardized labeling enables:
- Consistent results across experiments
- Reliable comparison between datasets
Multiplexing Capabilities
Chemical labeling allows simultaneous analysis of multiple samples, improving:
- Efficiency
- Throughput
- Data consistency
Compatibility with Complex Biological Samples
Labeled peptides perform well in:
- Plasma and serum
- Tissue extracts
- Cell lysates
Applications Across Research and Industry
Biomarker Discovery
Quantitative peptide analysis enables identification and validation of:
- Disease-specific biomarkers
- Prognostic indicators
- Therapeutic targets
Drug Development and Pharmacokinetics
Labeled peptides are used to:
- Monitor drug absorption and distribution
- Study metabolic pathways
- Evaluate drug efficacy
Proteomics and Systems Biology
Large-scale studies rely on labeled peptides to:
- Quantify protein expression
- Analyze signaling pathways
- Understand biological networks
Clinical Diagnostics
Quantitative peptide assays are increasingly applied in:
- Oncology diagnostics
- Metabolic disease monitoring
- Precision medicine
Challenges in Quantitative Peptide Labeling
Despite significant progress, challenges remain.
Cost and Accessibility
- Stable isotope-labeled peptides are expensive
- Multiplex labeling increases experimental cost
Sample Preparation Complexity
- Inconsistent labeling efficiency can affect accuracy
- Requires careful optimization
Instrumentation Requirements
- Advanced MS platforms are necessary
- Requires specialized expertise
Data Analysis Complexity
- Large datasets require advanced bioinformatics tools
- Interpretation can be computationally intensive
Best Practices for Reliable Quantitative Analysis
To ensure high-quality results:
✔ Use high-purity labeled standards
Reduces variability and improves calibration
✔ Optimize sample preparation
Ensure consistent labeling and handling
✔ Validate experimental methods
Confirm reproducibility across replicates
✔ Combine multiple analytical approaches
Integrate targeted and global analysis strategies
Future Trends in Quantitative Peptide Analysis
Integration with Artificial Intelligence
AI and machine learning are improving:
- Data processing
- Pattern recognition
- Predictive modeling
Ultra-Sensitive Detection
Advances in MS aim to enable:
- Single-cell proteomics
- Detection of ultra-low abundance peptides
Personalized Medicine
Quantitative peptide profiling supports:
- Patient-specific diagnostics
- Tailored therapeutic strategies
How LinkPeptide Supports Quantitative Peptide Research
At LinkPeptide, we provide advanced solutions tailored to quantitative peptide workflows, including:
- Stable isotope-labeled peptide synthesis
- Custom peptide standards for LC-MS/MS
- High-purity peptides with analytical validation
- Design optimization for quantitative accuracy
Our expertise enables researchers to achieve reliable, reproducible, and scalable peptide quantification across diverse applications.
Conclusion
Labeled peptides have revolutionized quantitative peptide analysis, providing the precision and reliability required for modern biological research. Through innovations in isotope labeling, chemical tagging, and mass spectrometry, researchers can now achieve highly accurate measurements even in complex biological systems.
As technology continues to evolve, labeled peptides will remain at the forefront of advances in drug discovery, clinical diagnostics, and systems biology, enabling deeper insights into molecular processes and accelerating scientific innovation.
Reference
Zhang, R., Sioma, C. S., Wang, S., & Regnier, F. E. (2001). Fractionation of isotopically labeled peptides in quantitative proteomics. Analytical chemistry, 73(21), 5142-5149. https://doi.org/10.1021/ac010583a
Richards, D. P., Sojo, L. E., & Keller, B. O. (2007). Quantitative analysis with modern bioanalytical mass spectrometry and stable isotope labeling. Journal of Labelled Compounds and Radiopharmaceuticals: The Official Journal of the International Isotope Society, 50(11‐12), 1124-1136. https://doi.org/10.1002/jlcr.1392
Treumann, A., & Thiede, B. (2010). Isobaric protein and peptide quantification: perspectives and issues. Expert review of proteomics, 7(5), 647-653. https://doi.org/10.1586/epr.10.29
Sauer, C. S., Phetsanthad, A., Riusech, O. L., & Li, L. (2021). Developing mass spectrometry for the quantitative analysis of neuropeptides. Expert review of proteomics, 18(7), 607-621. https://doi.org/10.1080/14789450.2021.1967146
 LinkPeptide
LinkPeptide