Selecting Peptide Substrates for Enzyme Assays: Principles, Design Strategies, and Analytical Considerations
Introduction: The Central Role of Peptide Substrates in Enzymology
Enzyme assays are indispensable tools in biochemical and biomedical research, underpinning studies in enzyme kinetics, drug discovery, signal transduction, and disease mechanism elucidation. The accuracy and interpretability of these assays depend critically on the choice of substrate. Among available options, peptide substrates have become particularly valuable due to their ability to replicate key features of natural protein targets while offering superior flexibility in design and modification.
Unlike full-length proteins, peptide substrates provide a simplified and controllable system for probing enzyme specificity and catalytic efficiency. However, this simplification introduces its own challenges: the selected peptide must retain sufficient structural and sequence information to faithfully represent the native substrate while remaining compatible with analytical detection methods.
A rational approach to peptide substrate selection is therefore essential. This involves integrating knowledge of enzyme specificity, structural biology, kinetics, and assay design to ensure that the resulting system produces reliable and biologically meaningful data.
Peptide Substrates: Definition and Functional Role
Peptide substrates are short amino acid sequences engineered to be recognized and processed by enzymes. They are widely used in assays involving:
- Proteases, which cleave peptide bonds
- Kinases, which phosphorylate specific residues
- Phosphatases, which remove phosphate groups
- Transferases, which catalyze group transfer reactions
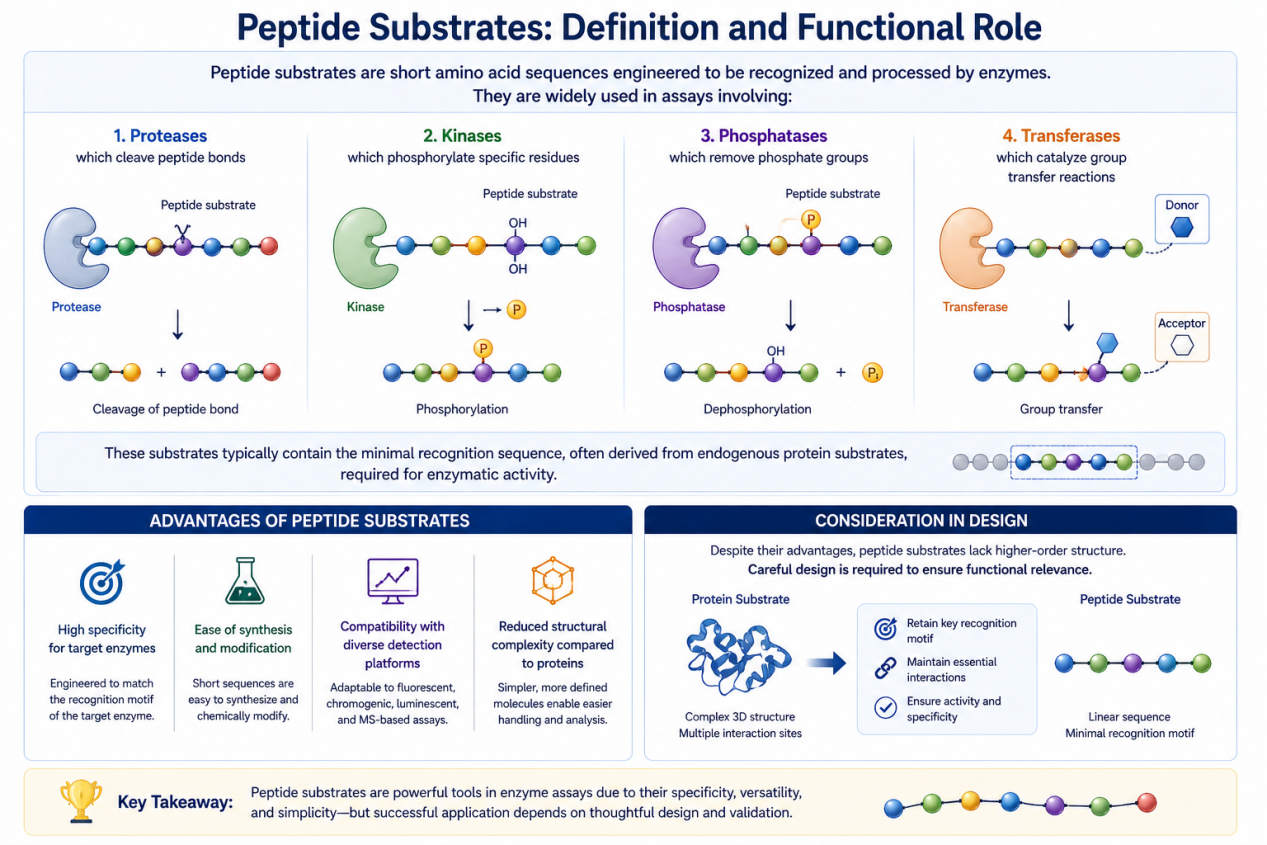
These substrates typically contain the minimal recognition sequence, often derived from endogenous protein substrates, required for enzymatic activity.
Advantages of Peptide Substrates
- High specificity for target enzymes
- Ease of synthesis and modification
- Compatibility with diverse detection platforms
- Reduced structural complexity compared to proteins
Despite these advantages, the absence of higher-order structure means that careful design is required to ensure functional relevance.
Molecular Basis of Enzyme–Substrate Recognition
Sequence Determinants and Positional Nomenclature
Enzyme specificity is governed by interactions between the substrate and defined binding pockets within the enzyme active site. These interactions are commonly described using the Schechter–Berger nomenclature, where:
- P1, P2, P3… represent residues N-terminal to the cleavage or modification site
- P1’, P2’, P3’… represent residues C-terminal to the site
Correspondingly, the enzyme binding sites are designated S1, S2, S3… and S1’, S2’, S3’.
The compatibility between substrate residues and enzyme subsites determines:
- Binding affinity
- Catalytic efficiency
- Substrate selectivity
Even single amino acid substitutions at critical positions can significantly alter enzyme activity, underscoring the importance of precise sequence design.
Key Criteria for Selecting Peptide Substrates
1. Sequence Specificity and Biological Relevance
The primary consideration is that the peptide sequence must reflect the enzyme’s natural substrate preference.
Approaches:
- Derivation from known physiological substrates
- Sequence alignment of protein targets
- Screening using peptide libraries
Maintaining biological relevance ensures that assay results can be extrapolated to in vivo systems.
2. Kinetic Suitability
A well-designed substrate must enable accurate kinetic analysis.
Key parameters include:
- Km (Michaelis constant): reflects substrate affinity
- kcat (turnover number): reflects catalytic efficiency
- kcat/Km: overall catalytic performance
An ideal substrate should exhibit:
- Moderate Km (to allow measurable binding)
- Sufficient turnover for detectable signal generation
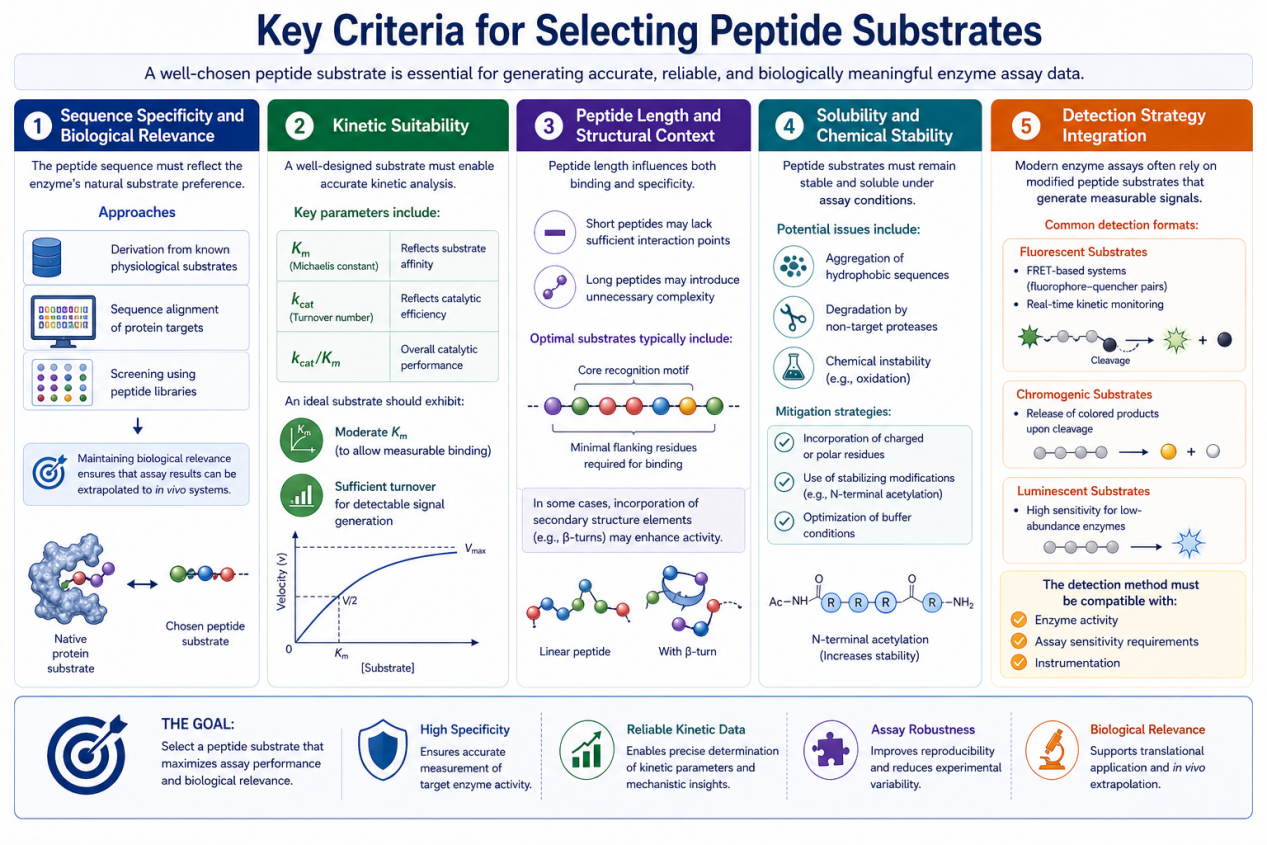
3. Peptide Length and Structural Context
Peptide length influences both binding and specificity.
- Short peptides may lack sufficient interaction points
- Long peptides may introduce unnecessary complexity
Optimal substrates typically include:
- Core recognition motif
- Minimal flanking residues required for binding
In some cases, incorporation of secondary structure elements (e.g., β-turns) may enhance activity.
4. Solubility and Chemical Stability
Peptide substrates must remain stable and soluble under assay conditions.
Potential issues include:
- Aggregation of hydrophobic sequences
- Degradation by non-target proteases
- Chemical instability (e.g., oxidation)
Mitigation strategies:
- Incorporation of charged or polar residues
- Use of stabilizing modifications (e.g., N-terminal acetylation)
- Optimization of buffer conditions
5. Detection Strategy Integration
Modern enzyme assays often rely on modified peptide substrates that generate measurable signals.
Common detection formats:
Fluorescent Substrates
- FRET-based systems (fluorophore–quencher pairs)
- Real-time kinetic monitoring
Chromogenic Substrates
- Release of colored products upon cleavage
Luminescent Substrates
- High sensitivity for low-abundance enzymes
The detection method must be compatible with:
- Enzyme activity
- Assay sensitivity requirements
- Instrumentation
Designing Substrates for Specific Enzyme Classes
Proteases
Proteases are among the most extensively studied enzymes using peptide substrates.
Design considerations:
- Accurate cleavage site identification
- Optimization of P1 and adjacent residues
- Avoidance of secondary cleavage sites
Advanced designs include quenched fluorescent substrates, which enable continuous monitoring of enzymatic activity.
Kinases
Kinase substrates require precise sequence motifs surrounding the phosphorylation site.
Key features:
- Presence of Ser, Thr, or Tyr residues
- Flanking residues that enhance recognition
Detection often involves:
- Radiolabeled phosphate incorporation
- Phospho-specific antibodies
- Fluorescent probes
Phosphatases
Phosphatase assays typically use pre-phosphorylated peptides.
Critical considerations:
- Stability of the phosphate group
- Sensitivity of detection method
Transferases and Other Enzymes
Substrate design must account for:
- Specific donor and acceptor groups
- Reaction mechanism
These assays often require specialized chemical modifications.
Optimization Strategies
Structure–Activity Relationship (SAR) Analysis
Systematic modification of peptide sequences allows identification of:
- Essential residues
- Optimal sequence length
- Key binding interactions
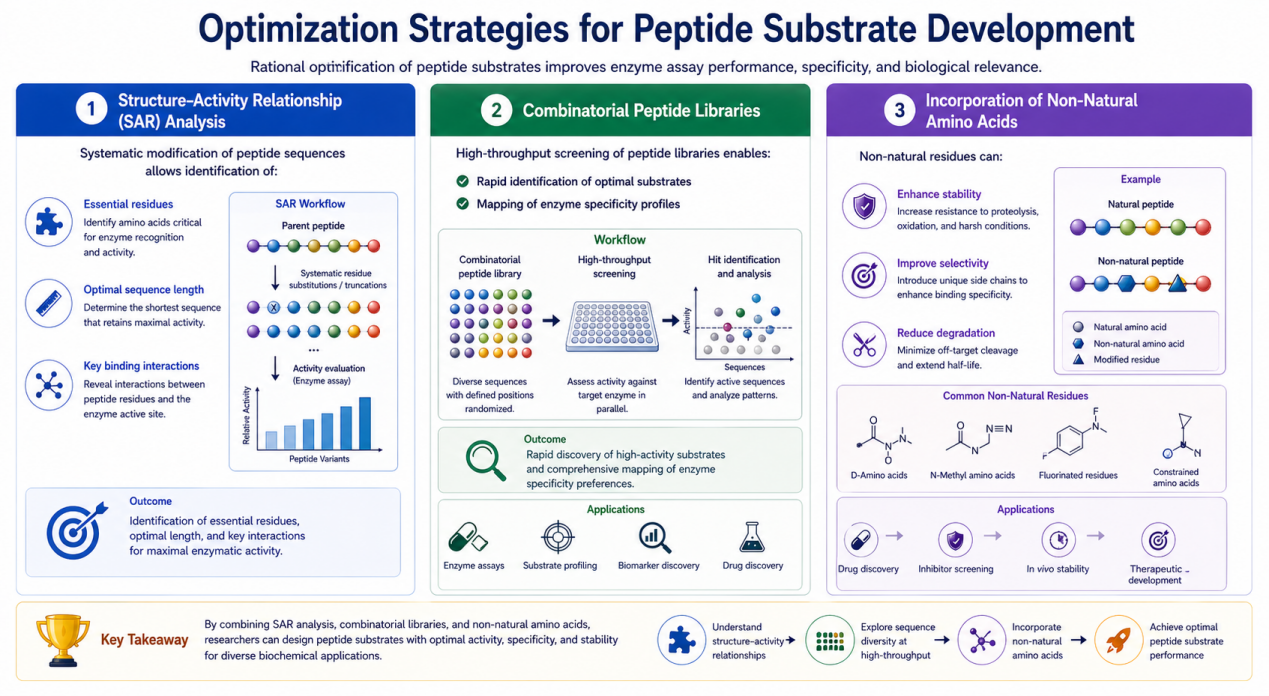
Combinatorial Peptide Libraries
High-throughput screening of peptide libraries enables:
- Rapid identification of optimal substrates
- Mapping of enzyme specificity profiles
Incorporation of Non-Natural Amino Acids
Non-natural residues can:
- Enhance stability
- Improve selectivity
- Reduce degradation
This is particularly useful in drug discovery and inhibitor screening.
Common Challenges in Peptide Substrate Design
Limited Specificity
Cross-reactivity with related enzymes can compromise assay accuracy.
Solution: Refine sequence and validate specificity experimentally.
Signal Limitations
Weak signal output reduces sensitivity.
Solution: Optimize labeling strategy and detection method.
Substrate Inhibition
High substrate concentrations may inhibit enzyme activity.
Solution: Perform concentration optimization and kinetic analysis.
Lack of Structural Context
Peptides may not fully replicate protein substrates.
Solution: Extend sequence or incorporate structural constraints.
Practical Considerations for Experimental Design
To ensure robust assay performance:
- Validate substrate specificity using control enzymes
- Optimize buffer composition and pH
- Confirm linearity of reaction rates
- Include appropriate positive and negative controls
- Perform replicate measurements to assess reproducibility
Applications of Peptide Substrates
Peptide substrates are widely used in:
- Drug discovery: screening enzyme inhibitors
- Cancer research: studying protease activity
- Signal transduction studies: kinase and phosphatase assays
- Infectious disease research: viral enzyme characterization
- Diagnostics: enzyme-based detection systems
Emerging Trends in Peptide Substrate Design
Recent developments include:
Smart Substrates
Activatable probes that respond to enzymatic activity
Multiplexed Assays
Simultaneous analysis of multiple enzymes
Integration with Mass Spectrometry
High-throughput quantitative analysis
Computational Design
Machine learning for predicting optimal substrates
These advances are driving more precise and scalable enzymatic studies.
How LinkPeptide Supports Peptide Substrate Development
At LinkPeptide, we provide comprehensive solutions for enzyme assay development, including:
- Custom peptide substrate design
- Fluorescent and labeled peptide synthesis
- Sequence optimization based on enzyme specificity
- High-purity peptides with analytical validation (HPLC, MS)
Our services enable researchers to develop high-performance peptide substrates tailored to their experimental systems.
Conclusion
The selection of peptide substrates is a critical determinant of enzyme assay performance. A rigorous approach that integrates sequence specificity, kinetic considerations, structural design, and detection strategy is essential for generating reliable and meaningful data.
With advances in peptide engineering and analytical technologies, researchers now have unprecedented capabilities to design substrates that closely mimic biological systems while offering superior experimental control.
As enzymology continues to evolve, well-designed peptide substrates will remain central to advancing our understanding of biochemical processes and therapeutic development.
Reference
Kemp, B. E., & Pearson, R. B. (1991). [10] Design and use of peptide substrates for protein kinases. Methods in enzymology, 200, 121-134.
https://doi.org/10.1016/0076-6879(91)00134-I
Fairlie, D. P., Tyndall, J. D., Reid, R. C., Wong, A. K., Abbenante, G., Scanlon, M. J., … & Burkett, B. A. (2000). Conformational selection of inhibitors and substrates by proteolytic enzymes: implications for drug design and polypeptide processing. Journal of Medicinal Chemistry, 43(7), 1271-1281.
https://doi.org/10.1021/jm990315t
Szymczak, L. C., Kuo, H. Y., & Mrksich, M. (2017). Peptide arrays: development and application. Analytical chemistry, 90(1), 266.
https://doi.org/10.1021/acs.analchem.7b04380
Li-Chan, E. C. (2015). Bioactive peptides and protein hydrolysates: research trends and challenges for application as nutraceuticals and functional food ingredients. Current Opinion in Food Science, 1, 28-37.
 LinkPeptide
LinkPeptide