Retro-Inverted Peptides: A New Design Rule for Precision Antimycobacterial Therapy
Abstract
Tuberculosis remains a global health emergency, exacerbated by the rapid rise of drug-resistant Mycobacterium tuberculosis and the inherent limitations of conventional antibiotics. Host defense peptides offer a promising alternative due to their membrane-targeting mechanisms, but their clinical potential has been constrained by poor stability and inconsistent activity. Recent research published in Nature Communications challenges this paradigm by demonstrating that peptide retro-inversion—a chemical modification traditionally used to enhance proteolytic resistance—can markedly improve both the potency and specificity of antimicrobial peptides against mycobacteria. Using the mycomembrane-targeting peptide MAD1 as a model, the study shows that fully retro-inverted analogues achieve order-of-magnitude gains in antimycobacterial activity while simultaneously reducing host toxicity. Mechanistic analyses reveal that these effects arise not from stability alone, but from altered peptide conformational rigidity and enhanced assembly within the unique mycobacterial outer membrane. Together, these findings position retro-inversion as a powerful and underappreciated design strategy for engineering next-generation, narrow-spectrum antimycobacterial peptides.
Tuberculosis at a Breaking Point: Why Conventional Antibiotics Are Falling Short
Tuberculosis (TB) remains one of the world’s most devastating infectious diseases, causing more than a million deaths each year despite decades of medical progress. While effective antibiotics exist, the global rise of multidrug-resistant and extensively drug-resistant Mycobacterium tuberculosis (Mtb) has exposed the fragility of current treatment strategies. Lengthy therapy regimens, significant side effects, and inconsistent patient adherence further compound the problem, allowing resistant strains to emerge and spread.
At the heart of this challenge lies the unique biology of the mycobacterial pathogen. Unlike most bacteria, M. tuberculosis is protected by a highly specialized, lipid-rich cell envelope dominated by long-chain mycolic acids. This so-called mycomembrane is exceptionally thick, rigid, and impermeable, acting as a formidable barrier to many small-molecule antibiotics. Even drugs that successfully reach their intracellular targets often exhibit slow kill kinetics and limited efficacy against non-replicating or persistent bacterial populations, enabling Mtb to survive prolonged therapeutic pressure.
Host defense peptides (HDPs) have gained attention as promising alternatives to conventional antibiotics because they operate through fundamentally different mechanisms. Rather than targeting a single enzyme or metabolic pathway, HDPs typically disrupt bacterial membranes and envelope integrity, a mode of action that is inherently difficult for pathogens to evade through mutation. Yet, despite their biological appeal, natural peptides have struggled to translate clinically due to poor proteolytic stability, potential host toxicity, and unpredictable performance in complex biological environments.
These shortcomings have prompted a shift in focus: from discovering new antimicrobial peptides to rationally engineering peptides with optimized stability, selectivity, and function. In this context, innovative design strategies are becoming essential to unlocking the therapeutic potential of peptide-based antimicrobials against tuberculosis.
Retro-Inversion Reconsidered: Challenging a Long-Standing Assumption in Peptide Design
One of the most widely used strategies to improve peptide drug stability is retro-inversion, a chemical modification in which the peptide sequence is reversed and composed entirely of D-amino acids. This approach preserves the relative orientation of side chains while rendering the peptide largely resistant to proteolytic degradation. For decades, retro-inversion has been viewed primarily as a practical workaround for one of peptide therapeutics’ greatest weaknesses: rapid enzymatic breakdown in biological environments.
However, experience has shown that the benefits of retro-inversion often come at a cost. Numerous studies across antibacterial, antiviral, and signaling peptides have reported that retro-inverted analogues lose biological potency, adopt altered secondary structures, or display impaired self-assembly. Changes in backbone direction and terminal group presentation can subtly but critically disrupt how peptides interact with membranes or protein targets. As a result, the prevailing assumption in peptide science has been clear: retro-inversion improves stability, but rarely improves—and often diminishes—biological function.
The study at the center of this discussion challenges that assumption in a striking and unexpected way. Rather than weakening antimicrobial activity, retro-inversion was shown to dramatically enhance both potency and specificity against mycobacteria. This effect was not marginal; in some cases, retro-inverted peptides exhibited order-of-magnitude improvements in activity compared with their native counterparts. Importantly, these gains were largely confined to mycobacterial species, suggesting that retro-inversion does more than protect peptides from degradation—it fundamentally alters how they engage with a uniquely structured bacterial target.
By reframing retro-inversion as a functional design strategy rather than a simple stability enhancement, this work opens the door to a new way of thinking about peptide engineering for difficult-to-treat pathogens like Mycobacterium tuberculosis.
When Peptide Chemistry Defies Expectations: Enhanced Potency and Selectivity Against Mycobacteria
To test whether retro-inversion could meaningfully alter antimycobacterial activity, the authors focused on a previously developed host defense peptide known as MAD1, which was originally designed to disrupt the mycobacterial outer membrane through biomimicry. MAD1 already displayed measurable activity against Mycobacterium tuberculosis, making it an ideal scaffold for systematic modification. The researchers generated three variants—one with reversed sequence direction, one composed of D-amino acids, and one combining both features to create a fully retro-inverted peptide, termed MAD1-RI.
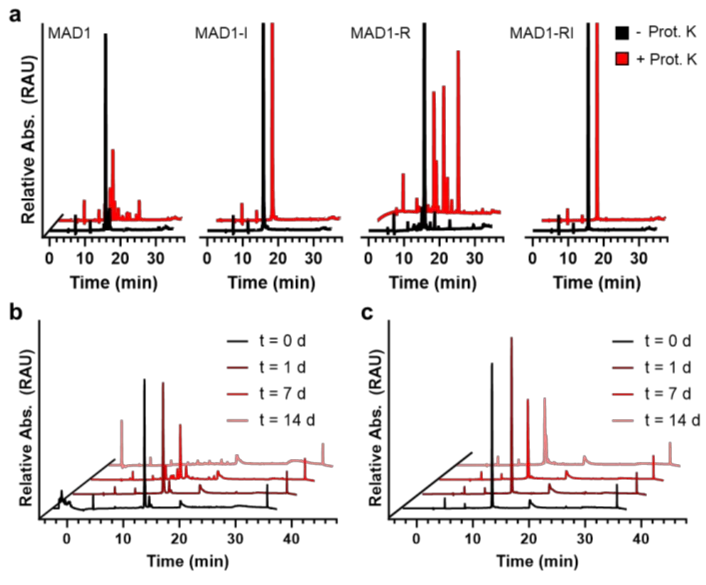
Figure 1. Proteolytic stability of MAD1 analogues. (a) Highpressure liquid chromatograms of the indicated peptide (100 μM) following a 1 hour incubation in the absence (black) and presence (red) of proteinase K. (b,c) Stability of MAD1 (b) and MAD1-RI (c) over two weeks of incubation in the proteolytic Mtb (mc2 6230) supernatant. Representative spectra shown from n = 3 technical replicates. Source data are provided as a Source Data file.
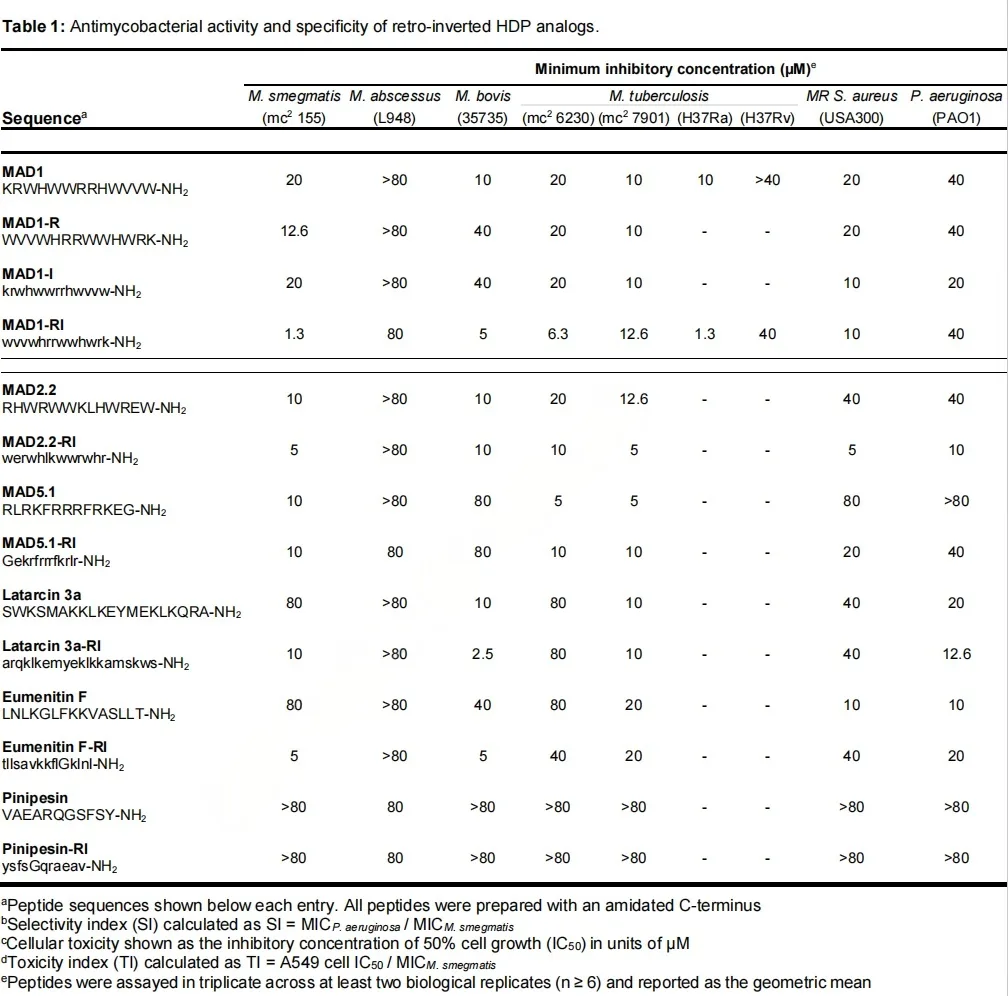
The results were striking. While sequence reversal or chirality inversion alone produced little change in antimicrobial potency, the fully retro-inverted MAD1-RI showed a dramatic improvement in activity against multiple mycobacterial species. Minimum inhibitory concentrations dropped into the low micromolar range for several M. tuberculosis strains, representing up to an order-of-magnitude enhancement compared with the native peptide. Notably, this increased potency extended to drug-resistant clinical isolates, highlighting the translational relevance of the finding.
Equally important was what did not change. When tested against non-mycobacterial pathogens such as Staphylococcus aureus and Pseudomonas aeruginosa, MAD1-RI did not exhibit a comparable increase in activity. This resulted in a substantial rise in mycobacterial selectivity, transforming MAD1-RI from a moderately active antimicrobial into a pathogen-focused agent. At the same time, retro-inversion markedly reduced toxicity toward human lung epithelial cells, yielding a greatly improved therapeutic index.
To assess whether this phenomenon was unique to MAD1, the authors extended their analysis to several unrelated antimicrobial peptides. In multiple cases, retro-inverted analogues again showed enhanced antimycobacterial potency, suggesting that this effect reflects a broader and previously underappreciated design principle rather than an isolated exception.
Cracking the Mycomembrane: How Retro-Inverted Peptides Achieve Superior Activity
At first glance, the enhanced activity of retro-inverted peptides might be attributed to their well-known resistance to proteolytic degradation. Indeed, the retro-inverted MAD1-RI peptide was highly stable in the presence of bacterial, serum, and broad-spectrum proteases. However, careful comparison of activity and stability revealed a critical insight: protease resistance alone could not explain the dramatic gains in antimycobacterial potency. Peptides that were protease-stable but not fully retro-inverted did not show comparable improvements in activity, indicating that a different mechanism was at play.
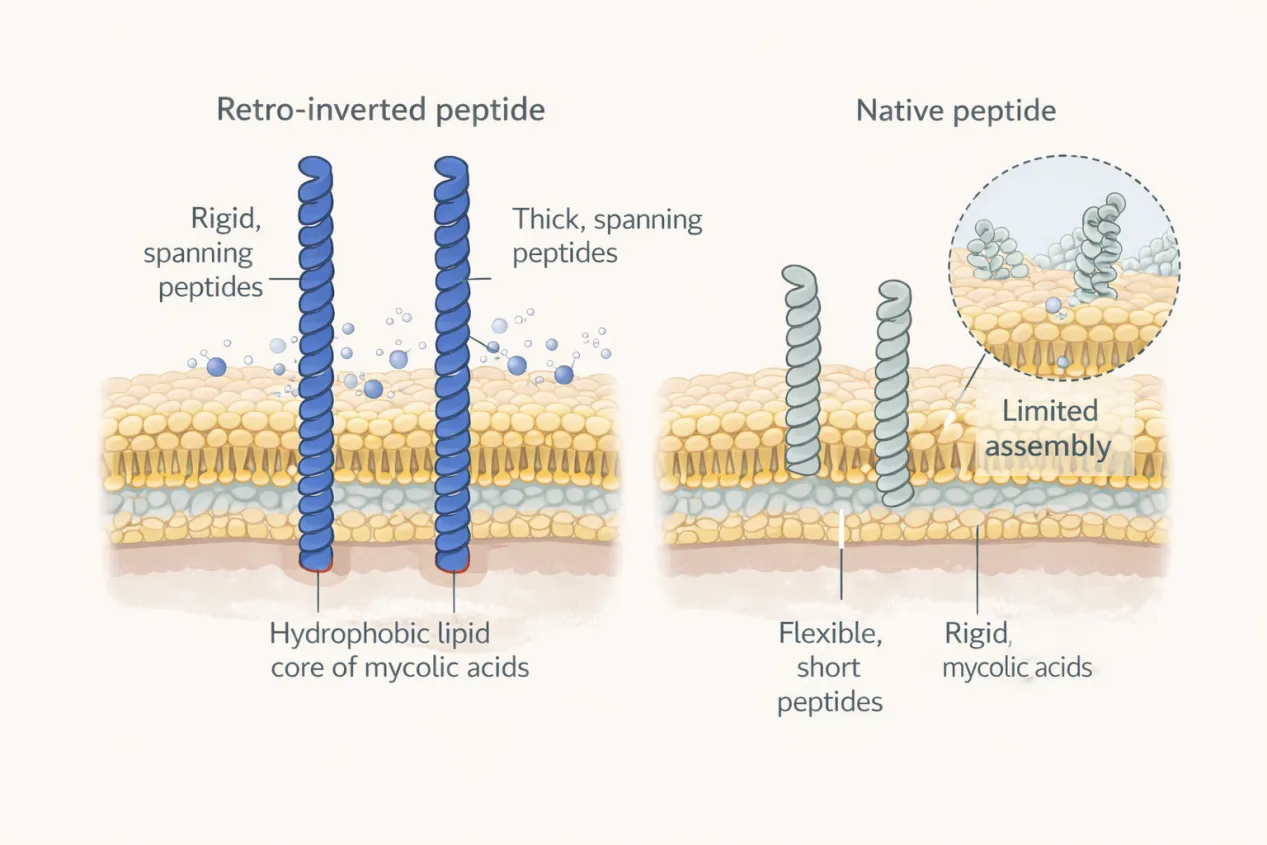
The answer lies in the unique physical properties of the mycobacterial cell envelope. The mycomembrane is unusually thick, highly ordered, and enriched in long-chain mycolic acids, creating a hydrophobic and rigid barrier unlike the membranes of most bacteria. Through an extensive series of biophysical experiments, the authors demonstrated that retro-inversion fundamentally alters how peptides interact with this environment.
Circular dichroism, small-angle X-ray scattering, and analytical ultracentrifugation revealed that MAD1-RI adopts a more conformationally rigid and thermodynamically stable structure than its native counterpart. Unlike MAD1, which readily loses structural order in response to changes in salt concentration, temperature, or detergent exposure, MAD1-RI maintains its secondary structure across a wide range of conditions. This rigidity appears to lower the energetic cost of membrane insertion, allowing the peptide to more readily engage and assemble within the mycomembrane.
Microscopy studies further showed that MAD1-RI accumulates at the mycobacterial surface and forms organized nano-assemblies that disrupt membrane integrity. Structural modeling suggested a head-to-tail dimerization mode for the retro-inverted peptide, producing assemblies capable of spanning the full thickness of the mycomembrane. This contrasts with the more limited assemblies formed by the native peptide and provides a plausible explanation for the enhanced membrane permeabilization observed experimentally.
Together, these findings point to altered mycomembrane thermodynamics and peptide self-assembly, rather than simple enzymatic stability, as the central drivers of retro-inversion–mediated antimycobacterial activity.
Beyond Stability: What Retro-Inversion Means for the Future of Antimicrobial Peptide Engineering
The implications of these findings extend well beyond a single peptide or experimental system. By demonstrating that retro-inversion can enhance potency, specificity, and safety against Mycobacterium tuberculosis, this work reframes retro-inversion as a rational design principle rather than a purely defensive modification to improve stability. In the context of antimicrobial peptide development, this distinction is critical. It suggests that chemical modifications can be deliberately leveraged to exploit pathogen-specific membrane properties, enabling the design of narrow-spectrum agents that minimize collateral damage to the host microbiota.
From a therapeutic perspective, the study highlights several attractive features of retro-inverted antimycobacterial peptides. MAD1-RI not only shows rapid bactericidal activity against replicating Mtb but also displays synergy with clinically relevant antibiotics such as moxifloxacin and meropenem. This opens the door to combination therapies that could shorten treatment duration, reduce drug dosing, and limit the emergence of resistance. Notably, repeated exposure experiments failed to induce resistance to MAD1-RI, underscoring the high mutational barrier associated with membrane-targeting mechanisms.
More broadly, these results carry important lessons for the future of peptide engineering. As artificial intelligence and computational tools accelerate the discovery of novel antimicrobial sequences, retro-inversion offers a complementary chemical strategy to fine-tune function, selectivity, and robustness. For difficult-to-treat pathogens protected by complex envelopes, such as mycobacteria, this approach may prove especially powerful.
Ultimately, this study suggests that the next generation of peptide antibiotics will not emerge solely from biological inspiration, but from precise molecular engineering informed by membrane biophysics—with retro-inversion standing out as a surprisingly potent tool in that design arsenal.
Reference
Medina, S., Glossop, H., Gebretsadik, G., Sultana, S., Schacht, N., Yennawar, N., … & Baughn, A. (2025). Retro-Inversion Imparts Antimycobacterial Specificity to Host Defense Peptides.https://doi.org/10.1038/s41467-025-67162-0
Rima, M., Rima, M., Fajloun, Z., Sabatier, J. M., Bechinger, B., & Naas, T. (2021). Antimicrobial peptides: A potent alternative to antibiotics. Antibiotics, 10(9), 1095.https://doi.org/10.3390/antibiotics10091095
Santos-Júnior, C. D., Torres, M. D., Duan, Y., Del Río, Á. R., Schmidt, T. S., Chong, H., … & Coelho, L. P. (2024). Discovery of antimicrobial peptides in the global microbiome with machine learning. Cell, 187(14), 3761-3778.https://doi.org/10.1016/j.cell.2024.05.013
Geberetsadik, G., Inaizumi, A., Nishiyama, A., Yamaguchi, T., Hamamoto, H., Panthee, S., … & Matsumoto, S. (2022). Lysocin E targeting menaquinone in the membrane of Mycobacterium tuberculosis is a promising lead compound for antituberculosis drugs. Antimicrobial Agents and Chemotherapy, 66(9), e00171-22.https://doi.org/10.1128/aac.00171-22
 LinkPeptide
LinkPeptide